Emergence and global dissemination of host-specific Streptococcus agalactiae clones
- PMID: 20824105
- PMCID: PMC2932510
- DOI: 10.1128/mBio.00178-10
Emergence and global dissemination of host-specific Streptococcus agalactiae clones
Abstract
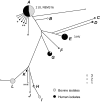
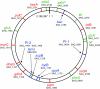
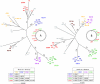
To examine the global diversity of Streptococcus agalactiae (group B streptococci [GBS]) and to elucidate the evolutionary processes that determine its population genetics structure and the reported changes in host tropism and infection epidemiology, we examined a collection of 238 bovine and human isolates from nine countries on five continents. Phylogenetic analysis based on the sequences of 15 housekeeping genes combined with patterns of virulence-associated traits identified a genetically heterogeneous core population from which virulent lineages occasionally emerge as a result of recombination affecting major segments of the genome. Such lineages, like clonal complex 17 (CC17) and two distinct clusters of CC23, are exclusively adapted to either humans or cattle and successfully spread globally. The recent emergence and expansion of the human-associated and highly virulent sequence type 17 (ST17) could conceivably account, in part, for the increased prevalence of neonatal GBS infections after 1960. The composite structure of the S. agalactiae genome invalidates phylogenetic inferences exclusively based on multilocus sequence typing (MLST) data and thereby the previously reported conclusion that the human-associated CC17 emerged from the bovine-associated CC67.
Figures

References
-
- Merl K., Abdulmawjood A., Lämmler C., Zschock M. 2003. Determination of epidemiological relationships of Streptococcus agalactiae isolated from bovine mastitis. FEMS Microbiol. Lett. 226:87–92 - PubMed
-
- Anthony B. F., Okada D. M. 1977. The emergence of group B streptococci in infections of the newborn infant. Annu. Rev. Med. 28:355–369 - PubMed
-
- Trijbels-Smeulders M. A., Kollee L. A., Adriaanse A. H., Kimpen J. L., Gerards L. J. 2004. Neonatal group B streptococcal infection: incidence and strategies for prevention in Europe. Pediatr. Infect. Dis. J. 23:172–173 - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

