Sequence- and structure-specific RNA processing by a CRISPR endonuclease
- PMID: 20829488
- PMCID: PMC3133607
- DOI: 10.1126/science.1192272
Sequence- and structure-specific RNA processing by a CRISPR endonuclease
Abstract
Many bacteria and archaea contain clustered regularly interspaced short palindromic repeats (CRISPRs) that confer resistance to invasive genetic elements. Central to this immune system is the production of CRISPR-derived RNAs (crRNAs) after transcription of the CRISPR locus. Here, we identify the endoribonuclease (Csy4) responsible for CRISPR transcript (pre-crRNA) processing in Pseudomonas aeruginosa. A 1.8 angstrom crystal structure of Csy4 bound to its cognate RNA reveals that Csy4 makes sequence-specific interactions in the major groove of the crRNA repeat stem-loop. Together with electrostatic contacts to the phosphate backbone, these enable Csy4 to bind selectively and cleave pre-crRNAs using phylogenetically conserved serine and histidine residues in the active site. The RNA recognition mechanism identified here explains sequence- and structure-specific processing by a large family of CRISPR-specific endoribonucleases.
Figures
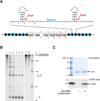


References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

