Glycosylation of pilin and nonpilin protein constructs by Pseudomonas aeruginosa 1244
- PMID: 20833803
- PMCID: PMC2976441
- DOI: 10.1128/JB.00007-10
Glycosylation of pilin and nonpilin protein constructs by Pseudomonas aeruginosa 1244
Abstract
PilO is an oligosaccharyl transferase (OTase) that catalyzes the O-glycosylation of Pseudomonas aeruginosa 1244 pilin by adding a single O-antigen repeating unit to the β carbon of the C-terminal residue (a serine). While PilO has an absolute requirement for Ser/Thr at this position, it is unclear if this enzyme must recognize other pilin features. To test this, pilin constructs containing peptide extensions terminating with serine were tested for the ability to support glycosylation. It was found that a 15-residue peptide, which had been modeled on the C-proximal region of strain 1244 pilin, served as a PilO substrate when it was expressed on either group II or group III pilins. In addition, adding a 3-residue extension culminating in serine to the C terminus of a group III pilin supported PilO activity. A protein fusion composed of strain 1244 pilin linked at its C terminus with Escherichia coli alkaline phosphatase (which, in turn, contained the above-mentioned 15 amino acids at its C terminus) was glycosylated by PilO. E. coli alkaline phosphatase lacking the pilin membrane anchor and containing the 15-residue peptide was also glycosylated by PilO. Addition of the 3-residue extension did not allow glycosylation of either of these constructs. Site-directed mutagenesis of strain 1244 pilin residues of the C-proximal region common to the group I proteins showed that this structure was not required for glycosylation. These experiments indicate that pilin common sequence is not required for glycosylation and show that nonpilin protein can be engineered to be a PilO substrate.
Figures






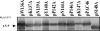
Similar articles
-
PilO of Pseudomonas aeruginosa 1244: subcellular location and domain assignment.Mol Microbiol. 2007 Dec;66(6):1444-58. doi: 10.1111/j.1365-2958.2007.06001.x. Epub 2007 Nov 13. Mol Microbiol. 2007. PMID: 18005110 Free PMC article.
-
Glycosylation of Pseudomonas aeruginosa 1244 pilin: glycan substrate specificity.Mol Microbiol. 2002 Oct;46(2):519-30. doi: 10.1046/j.1365-2958.2002.03171.x. Mol Microbiol. 2002. PMID: 12406226
-
Modification of Pseudomonas aeruginosa Pa5196 type IV Pilins at multiple sites with D-Araf by a novel GT-C family Arabinosyltransferase, TfpW.J Bacteriol. 2008 Nov;190(22):7464-78. doi: 10.1128/JB.01075-08. Epub 2008 Sep 19. J Bacteriol. 2008. PMID: 18805982 Free PMC article.
-
Glycosylation substrate specificity of Pseudomonas aeruginosa 1244 pilin.J Biol Chem. 2006 Jan 13;281(2):1128-36. doi: 10.1074/jbc.M510975200. Epub 2005 Nov 11. J Biol Chem. 2006. PMID: 16286455 Free PMC article.
-
Posttranslational processing of type IV prepilin and homologs by PilD of Pseudomonas aeruginosa.Methods Enzymol. 1994;235:527-40. doi: 10.1016/0076-6879(94)35168-6. Methods Enzymol. 1994. PMID: 8057924 Review.
Cited by
-
Synthetic Glycobiology: Parts, Systems, and Applications.ACS Synth Biol. 2020 Jul 17;9(7):1534-1562. doi: 10.1021/acssynbio.0c00210. Epub 2020 Jun 30. ACS Synth Biol. 2020. PMID: 32526139 Free PMC article. Review.
-
The type IV pili component PilO is a virulence determinant of Francisella novicida.PLoS One. 2022 Jan 25;17(1):e0261938. doi: 10.1371/journal.pone.0261938. eCollection 2022. PLoS One. 2022. PMID: 35077486 Free PMC article.
-
The sweet tooth of bacteria: common themes in bacterial glycoconjugates.Microbiol Mol Biol Rev. 2014 Sep;78(3):372-417. doi: 10.1128/MMBR.00007-14. Microbiol Mol Biol Rev. 2014. PMID: 25184559 Free PMC article. Review.
-
Glycoengineering bioconjugate vaccines, therapeutics, and diagnostics in E. coli.Glycobiology. 2019 Jul 1;29(7):519-529. doi: 10.1093/glycob/cwz031. Glycobiology. 2019. PMID: 30989179 Free PMC article. Review.
-
Construction, expression, purification and characterization of secretin domain of PilQ and triple PilA-related disulfide loop peptides fusion protein from Pseudomonas aeruginosa.Iran J Basic Med Sci. 2017 May;20(5):458-466. doi: 10.22038/IJBMS.2017.8667. Iran J Basic Med Sci. 2017. PMID: 28656079 Free PMC article.
References
-
- Aas, F. E., W. Egge-Jacobsen, H. C. Winther-Larsen, C. Lovold, P. G. Hitchen, A. Dell, and M. Koomey. 2006. Neisseria gonorrhoeae type IV pili undergo multisite hierarchical modifications with phosphoethanolamine and phosphocholine requiring an enzyme structurally related to lipopolysaccharide phosphoethanolamine transferases. J. Biol. Chem. 281:27712-27723. - PubMed
-
- Bolivar, F., and K. Backman. 1979. Plasmids of Escherichia coli as cloning vectors. Methods Enzymol. 68:245-267. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

