Computational modeling of mammalian signaling networks
- PMID: 20836022
- PMCID: PMC3105527
- DOI: 10.1002/wsbm.52
Computational modeling of mammalian signaling networks
Abstract
One of the most exciting developments in signal transduction research has been the proliferation of studies in which a biological discovery was initiated by computational modeling. In this study, we review the major efforts that enable such studies. First, we describe the experimental technologies that are generally used to identify the molecular components and interactions in, and dynamic behavior exhibited by, a network of interest. Next, we review the mathematical approaches that are used to model signaling network behavior. Finally, we focus on three specific instances of 'model-driven discovery': cases in which computational modeling of a signaling network has led to new insights that have been verified experimentally.
Figures
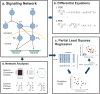

Similar articles
-
Modeling of spatially-restricted intracellular signaling.Wiley Interdiscip Rev Syst Biol Med. 2012 Jan-Feb;4(1):103-15. doi: 10.1002/wsbm.155. Epub 2011 Jul 15. Wiley Interdiscip Rev Syst Biol Med. 2012. PMID: 21766466 Free PMC article. Review.
-
Computational multiscale modeling of embryo development.Curr Opin Genet Dev. 2012 Dec;22(6):613-8. doi: 10.1016/j.gde.2012.08.006. Epub 2012 Sep 6. Curr Opin Genet Dev. 2012. PMID: 22959149 Review.
-
A data-driven computational model of the ErbB receptor signaling network.Conf Proc IEEE Eng Med Biol Soc. 2006;2006:53-4. doi: 10.1109/IEMBS.2006.259754. Conf Proc IEEE Eng Med Biol Soc. 2006. PMID: 17946779
-
Computational approaches for analyzing information flow in biological networks.Sci Signal. 2012 Apr 17;5(220):re1. doi: 10.1126/scisignal.2002961. Sci Signal. 2012. PMID: 22510471
-
Network modeling of signal transduction: establishing the global view.Bioessays. 2008 Nov;30(11-12):1110-25. doi: 10.1002/bies.20834. Bioessays. 2008. PMID: 18937364 Review.
Cited by
-
A forecast for large-scale, predictive biology: Lessons from meteorology.Cell Syst. 2021 Jun 16;12(6):488-496. doi: 10.1016/j.cels.2021.05.014. Cell Syst. 2021. PMID: 34139161 Free PMC article. Review.
-
Coherent functional modules improve transcription factor target identification, cooperativity prediction, and disease association.PLoS Genet. 2014 Feb 6;10(2):e1004122. doi: 10.1371/journal.pgen.1004122. eCollection 2014 Feb. PLoS Genet. 2014. PMID: 24516403 Free PMC article.
-
Signaling network prediction by the Ontology Fingerprint enhanced Bayesian network.BMC Syst Biol. 2012;6 Suppl 3(Suppl 3):S3. doi: 10.1186/1752-0509-6-S3-S3. Epub 2012 Dec 17. BMC Syst Biol. 2012. PMID: 23282239 Free PMC article.
-
Visualization of the spatial and temporal dynamics of MAPK signaling using fluorescence imaging techniques.J Physiol Sci. 2015 Jan;65(1):37-49. doi: 10.1007/s12576-014-0332-9. Epub 2014 Aug 22. J Physiol Sci. 2015. PMID: 25145828 Free PMC article. Review.
-
Histamine-induced biphasic activation of RhoA allows for persistent RhoA signaling.PLoS Biol. 2020 Sep 3;18(9):e3000866. doi: 10.1371/journal.pbio.3000866. eCollection 2020 Sep. PLoS Biol. 2020. PMID: 32881857 Free PMC article.
References
-
- Martin GS. Cell signaling and cancer. Cancer Cell. 2003;4(3):167–74. - PubMed
-
- Irish JM, Kotecha N, Nolan GP. Mapping normal and cancer cell signalling networks: towards single-cell proteomics. Nat Rev Cancer. 2006;6(2):146–55. - PubMed
-
- Fields S, Song O. A novel genetic system to detect protein-protein interactions. Nature. 1989;340(6230):245–6. - PubMed
-
- Parrish JR, Gulyas KD, Finley RL., Jr Yeast two-hybrid contributions to interactome mapping. Curr Opin Biotechnol. 2006;17(4):387–93. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

