Evolution of the Drosophila feminizing switch gene Sex-lethal
- PMID: 20837995
- PMCID: PMC2998314
- DOI: 10.1534/genetics.110.121202
Evolution of the Drosophila feminizing switch gene Sex-lethal
Abstract
In Drosophila melanogaster, the gene Sex-lethal (Sxl) controls all aspects of female development. Since melanogaster males lacking Sxl appear wild type, Sxl would seem to be functionally female specific. Nevertheless, in insects as diverse as honeybees and houseflies, Sxl seems not to determine sex or to be functionally female specific. Here we describe three lines of work that address the questions of how, when, and even whether the ancestor of melanogaster Sxl ever shed its non-female-specific functions. First, to test the hypothesis that the birth of Sxl's closest paralog allowed Sxl to lose essential ancestral non-female-specific functions, we determined the CG3056 null phenotype. That phenotype failed to support this hypothesis. Second, to define when Sxl might have lost ancestral non-female-specific functions, we isolated and characterized Sxl mutations in D. virilis, a species distant from melanogaster and notable for the large amount of Sxl protein expression in males. We found no change in Sxl regulation or functioning in the 40+ MY since these two species diverged. Finally, we discovered conserved non-sex-specific Sxl mRNAs containing a previously unknown, potentially translation-initiating exon, and we identified a conserved open reading frame starting in Sxl male-specific exon 3. We conclude that Drosophila Sxl may appear functionally female specific not because it lost non-female-specific functions, but because those functions are nonessential in the laboratory. The potential evolutionary relevance of these nonessential functions is discussed.
Figures



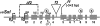



Similar articles
-
Evolution of the sex-lethal gene in insects and origin of the sex-determination system in Drosophila.J Mol Evol. 2014 Jan;78(1):50-65. doi: 10.1007/s00239-013-9599-3. Epub 2013 Nov 24. J Mol Evol. 2014. PMID: 24271947
-
Functional conservation of the sex-lethal sex determining promoter, Sxl-Pe, in Drosophila virilis.Dev Genes Evol. 2003 May;213(4):155-65. doi: 10.1007/s00427-003-0304-1. Epub 2003 Feb 12. Dev Genes Evol. 2003. PMID: 12739140
-
Transposon insertions causing constitutive Sex-lethal activity in Drosophila melanogaster affect Sxl sex-specific transcript splicing.Genetics. 1995 Feb;139(2):631-48. doi: 10.1093/genetics/139.2.631. Genetics. 1995. PMID: 7713421 Free PMC article.
-
RNA binding protein sex-lethal (Sxl) and control of Drosophila sex determination and dosage compensation.Microbiol Mol Biol Rev. 2003 Sep;67(3):343-59, table of contents. doi: 10.1128/MMBR.67.3.343-359.2003. Microbiol Mol Biol Rev. 2003. PMID: 12966139 Free PMC article. Review.
-
Sex determination in Drosophila: The view from the top.Fly (Austin). 2010 Jan-Mar;4(1):60-70. doi: 10.4161/fly.4.1.11277. Epub 2010 Jan 21. Fly (Austin). 2010. PMID: 20160499 Free PMC article. Review.
Cited by
-
Evolution of sexual development and sexual dimorphism in insects.Curr Opin Genet Dev. 2021 Aug;69:129-139. doi: 10.1016/j.gde.2021.02.011. Epub 2021 Apr 10. Curr Opin Genet Dev. 2021. PMID: 33848958 Free PMC article. Review.
-
Evolution of the sex-lethal gene in insects and origin of the sex-determination system in Drosophila.J Mol Evol. 2014 Jan;78(1):50-65. doi: 10.1007/s00239-013-9599-3. Epub 2013 Nov 24. J Mol Evol. 2014. PMID: 24271947
-
Drosophila Sister-of-Sex-lethal is a repressor of translation.RNA. 2018 Feb;24(2):149-158. doi: 10.1261/rna.063776.117. Epub 2017 Oct 31. RNA. 2018. PMID: 29089381 Free PMC article.
-
Influence of Wolbachia on host gene expression in an obligatory symbiosis.BMC Microbiol. 2012 Jan 18;12 Suppl 1(Suppl 1):S7. doi: 10.1186/1471-2180-12-S1-S7. BMC Microbiol. 2012. PMID: 22376153 Free PMC article.
-
Repeated Evolution of Asexuality Involves Convergent Gene Expression Changes.Mol Biol Evol. 2019 Feb 1;36(2):350-364. doi: 10.1093/molbev/msy217. Mol Biol Evol. 2019. PMID: 30445505 Free PMC article.
References
-
- Alexander, M. L., 1976. The Genetics and Biology of Drosophila virilis. Academic Press, London.
-
- Bopp, D., L. R. Bell, T. W. Cline and P. Schedl, 1991. Developmental distribution of female-specific Sex-lethal proteins in Drosophila melanogaster. Genes Dev. 5 403–415. - PubMed
-
- Bopp, D., G. Calhoun, J. I. Horabin, M. Samuels and P. Schedl, 1996. Sex-specific control of Sex-lethal is a conserved mechanism for sex determination in the genus Drosophila. Development 122 971–982. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

