A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis
- PMID: 20838596
- PMCID: PMC2936532
- DOI: 10.1371/journal.pgen.1001102
A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis
Abstract
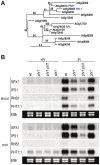
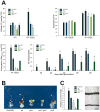
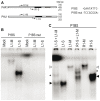
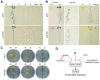
Plants respond to different stresses by inducing or repressing transcription of partially overlapping sets of genes. In Arabidopsis, the PHR1 transcription factor (TF) has an important role in the control of phosphate (Pi) starvation stress responses. Using transcriptomic analysis of Pi starvation in phr1, and phr1 phr1-like (phl1) mutants and in wild type plants, we show that PHR1 in conjunction with PHL1 controls most transcriptional activation and repression responses to phosphate starvation, regardless of the Pi starvation specificity of these responses. Induced genes are enriched in PHR1 binding sequences (P1BS) in their promoters, whereas repressed genes do not show such enrichment, suggesting that PHR1(-like) control of transcriptional repression responses is indirect. In agreement with this, transcriptomic analysis of a transgenic plant expressing PHR1 fused to the hormone ligand domain of the glucocorticoid receptor showed that PHR1 direct targets (i.e., displaying altered expression after GR:PHR1 activation by dexamethasone in the presence of cycloheximide) corresponded largely to Pi starvation-induced genes that are highly enriched in P1BS. A minimal promoter containing a multimerised P1BS recapitulates Pi starvation-specific responsiveness. Likewise, mutation of P1BS in the promoter of two Pi starvation-responsive genes impaired their responsiveness to Pi starvation, but not to other stress types. Phylogenetic footprinting confirmed the importance of P1BS and PHR1 in Pi starvation responsiveness and indicated that P1BS acts in concert with other cis motifs. All together, our data show that PHR1 and PHL1 are partially redundant TF acting as central integrators of Pi starvation responses, both specific and generic. In addition, they indicate that transcriptional repression responses are an integral part of adaptive responses to stress.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Mol J, Jenkins G, Schäfer E, Weiss D. Signal perception, transduction, and gene expression involved in anthocyanin. Crit Rev Plant Sci. 1996;15:525–557.
-
- Lim PO, Kim HJ, Nam HG. Leaf senescence. Annu Rev Plant Biol. 2007;58:115–136. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

