New insights into the fructosyltransferase activity of Schwanniomyces occidentalis ß-fructofuranosidase, emerging from nonconventional codon usage and directed mutation
- PMID: 20851958
- PMCID: PMC2976189
- DOI: 10.1128/AEM.01614-10
New insights into the fructosyltransferase activity of Schwanniomyces occidentalis ß-fructofuranosidase, emerging from nonconventional codon usage and directed mutation
Abstract
Schwanniomyces occidentalis β-fructofuranosidase (Ffase) releases β-fructose from the nonreducing ends of β-fructans and synthesizes 6-kestose and 1-kestose, both considered prebiotic fructooligosaccharides. Analyzing the amino acid sequence of this protein revealed that it includes a serine instead of a leucine at position 196, caused by a nonuniversal decoding of the unique mRNA leucine codon CUG. Substitution of leucine for Ser196 dramatically lowers the apparent catalytic efficiency (k(cat)/K(m)) of the enzyme (approximately 1,000-fold), but surprisingly, its transferase activity is enhanced by almost 3-fold, as is the enzymes' specificity for 6-kestose synthesis. The influence of 6 Ffase residues on enzyme activity was analyzed on both the Leu196/Ser196 backgrounds (Trp47, Asn49, Asn52, Ser111, Lys181, and Pro232). Only N52S and P232V mutations improved the transferase activity of the wild-type enzyme (about 1.6-fold). Modeling the transfructosylation products into the active site, in combination with an analysis of the kinetics and transfructosylation reactions, defined a new region responsible for the transferase specificity of the enzyme.
Figures




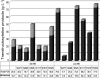

References
-
- Abarca, D., M. Fernández-Lobato, L. del Pozo, and A. Jiménez. 1991. Isolation of a new gene (SWA2) encoding an alpha amylase from Schwanniomyces occidentalis and its expression in Saccharomyces cerevisiae. FEBS Lett. 279:41-44. - PubMed
-
- Alberto, F., C. Bignon, G. Sulzenbacher, B. Henrissat, and M. Czjzek. 2004. The three-dimenisonal structure of invertase (β-fructosidase) from Thermotoga maritima reveals a bimodular arrangement and an evolutionary relationship between retaining and inverting glycosidases. J. Biol. Chem. 279:18903-18910. - PubMed
-
- Álvaro-Benito, M., M. de Abreu, L. Fernández-Arrojo, F. J. Plou, J. Jiménez-Barbero, A. Ballesteros, J. Polaina, and M. Fernández-Lobato. 2007. Characterization of a β-fructofuranosidase from Schwanniomyces occidentalis with transfructosylating activity yielding the prebiotic 6-kestose. J. Biotechnol. 132:75-81. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

