Stable, site-specific fluorescent tagging constructs optimized for burkholderia species
- PMID: 20851961
- PMCID: PMC2976199
- DOI: 10.1128/AEM.01188-10
Stable, site-specific fluorescent tagging constructs optimized for burkholderia species
Abstract
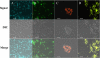
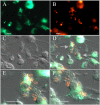
Several vectors that facilitate stable fluorescent labeling of Burkholderia pseudomallei and Burkholderia thailandensis were constructed. These vectors combined the effectiveness of the mini-Tn7 site-specific transposition system with fluorescent proteins optimized for Burkholderia spp., enabling bacterial tracking during cellular infection.
Figures


Similar articles
-
Engineering of tellurite-resistant genetic tools for single-copy chromosomal analysis of Burkholderia spp. and characterization of the Burkholderia thailandensis betBA operon.Appl Environ Microbiol. 2009 Jun;75(12):4015-27. doi: 10.1128/AEM.02733-08. Epub 2009 Apr 17. Appl Environ Microbiol. 2009. PMID: 19376905 Free PMC article.
-
Tn5/7-lux: a versatile tool for the identification and capture of promoters in gram-negative bacteria.BMC Microbiol. 2015 Feb 4;15(1):17. doi: 10.1186/s12866-015-0354-3. BMC Microbiol. 2015. PMID: 25648327 Free PMC article.
-
Sequence-defined transposon mutant library of Burkholderia thailandensis.mBio. 2013 Nov 5;4(6):e00604-13. doi: 10.1128/mBio.00604-13. mBio. 2013. PMID: 24194535 Free PMC article.
-
Molecular motors: single-molecule recordings made easy.Curr Biol. 2002 Mar 19;12(6):R203-5. doi: 10.1016/s0960-9822(02)00750-9. Curr Biol. 2002. PMID: 11909546 Review.
-
Green fluorescent protein (GFP) fusion constructs in gene therapy research.Histochem Cell Biol. 2001 Jan;115(1):59-65. doi: 10.1007/s004180000219. Histochem Cell Biol. 2001. PMID: 11219609 Review.
Cited by
-
The Burkholderia bcpAIOB genes define unique classes of two-partner secretion and contact dependent growth inhibition systems.PLoS Genet. 2012;8(8):e1002877. doi: 10.1371/journal.pgen.1002877. Epub 2012 Aug 9. PLoS Genet. 2012. PMID: 22912595 Free PMC article.
-
An improved recombination-based in vivo expression technology-like reporter system reveals differential cyaA gene activation in Bordetella species.Infect Immun. 2013 Apr;81(4):1295-305. doi: 10.1128/IAI.01445-12. Epub 2013 Feb 4. Infect Immun. 2013. PMID: 23381998 Free PMC article.
-
A virulence activator of a surface attachment protein in Burkholderia pseudomallei acts as a global regulator of other membrane-associated virulence factors.Front Microbiol. 2023 Jan 16;13:1063287. doi: 10.3389/fmicb.2022.1063287. eCollection 2022. Front Microbiol. 2023. PMID: 36726566 Free PMC article.
-
Structural diversity of Burkholderia pseudomallei lipopolysaccharides affects innate immune signaling.PLoS Negl Trop Dis. 2017 Apr 28;11(4):e0005571. doi: 10.1371/journal.pntd.0005571. eCollection 2017 Apr. PLoS Negl Trop Dis. 2017. PMID: 28453531 Free PMC article.
-
Imaging flow cytometry analysis of intracellular pathogens.Methods. 2017 Jan 1;112:91-104. doi: 10.1016/j.ymeth.2016.09.007. Epub 2016 Sep 15. Methods. 2017. PMID: 27642004 Free PMC article. Review.
References
-
- Biery, M. C., M. Lopata, and N. L. Craig. 2000. A minimal system for Tn7 transposition: the transposon-encoded proteins TnsA and TnsB can execute DNA breakage and joining reactions that generate circularized Tn7 species. J. Mol. Biol. 297:25-37. - PubMed
-
- Brett, P. J., D. DeShazer, and D. E. Woods. 1998. Burkholderia thailandensis sp. nov., description of Burkholderia pseudomallei-like species. Int. J. Syst. Bacteriol. 48:317-320. - PubMed
-
- Castle, L. A., D. L. Siehl, R. Gorton, P. A. Patten, Y. H. Chen, S. Bertain, H. Cho, N. Duck, J. Wong, D. Liu, and M. W. Lassner. 2004. Discovery and directed evolution of a glyphosate tolerance gene. Science 304:1151-1154. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

