Development of an environmental functional gene microarray for soil microbial communities
- PMID: 20851978
- PMCID: PMC2976231
- DOI: 10.1128/AEM.03108-09
Development of an environmental functional gene microarray for soil microbial communities
Abstract
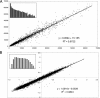
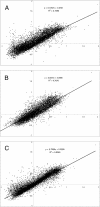
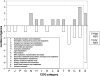
Functional attributes of microbial communities are difficult to study, and most current techniques rely on DNA- and rRNA-based profiling of taxa and genes, including microarrays containing sequences of known microorganisms. To quantify gene expression in environmental samples in a culture-independent manner, we constructed an environmental functional gene microarray (E-FGA) consisting of 13,056 mRNA-enriched anonymous microbial clones from diverse microbial communities to profile microbial gene transcripts. A new normalization method using internal spot standards was devised to overcome spotting and hybridization bias, enabling direct comparisons of microarrays. To evaluate potential applications of this metatranscriptomic approach for studying microbes in environmental samples, we tested the E-FGA by profiling the microbial activity of agricultural soils with a low or high flux of N₂O. A total of 109 genes displayed expression that differed significantly between soils with low and high N₂O emissions. We conclude that mRNA-based approaches such as the one presented here may complement existing techniques for assessing functional attributes of microbial communities.
Figures

Similar articles
-
Phylogenetic and Functional Diversity of Total (DNA) and Expressed (RNA) Bacterial Communities in Urban Green Infrastructure Bioswale Soils.Appl Environ Microbiol. 2017 Aug 1;83(16):e00287-17. doi: 10.1128/AEM.00287-17. Print 2017 Aug 15. Appl Environ Microbiol. 2017. PMID: 28576763 Free PMC article.
-
Direct detection of 16S rRNA in soil extracts by using oligonucleotide microarrays.Appl Environ Microbiol. 2001 Oct;67(10):4708-16. doi: 10.1128/AEM.67.10.4708-4716.2001. Appl Environ Microbiol. 2001. PMID: 11571176 Free PMC article.
-
Hybridization of genomic DNA to microarrays: a challenge for the analysis of environmental samples.J Microbiol Methods. 2007 May;69(2):242-8. doi: 10.1016/j.mimet.2006.11.007. Epub 2006 Dec 26. J Microbiol Methods. 2007. PMID: 17188770 Review.
-
Development of a common oligonucleotide reference standard for microarray data normalization and comparison across different microbial communities.Appl Environ Microbiol. 2010 Feb;76(4):1088-94. doi: 10.1128/AEM.02749-09. Epub 2009 Dec 28. Appl Environ Microbiol. 2010. PMID: 20038701 Free PMC article.
-
Microbial gene expression in soil: methods, applications and challenges.J Microbiol Methods. 2005 Oct;63(1):1-19. doi: 10.1016/j.mimet.2005.03.007. J Microbiol Methods. 2005. PMID: 15939495 Review.
Cited by
-
Linking microbial community structure to β-glucosidic function in soil aggregates.ISME J. 2013 Oct;7(10):2044-53. doi: 10.1038/ismej.2013.87. Epub 2013 May 30. ISME J. 2013. PMID: 23719152 Free PMC article.
-
Identification of active denitrifiers in rice paddy soil by DNA- and RNA-based analyses.Microbes Environ. 2012;27(4):456-61. doi: 10.1264/jsme2.me12076. Epub 2012 Sep 5. Microbes Environ. 2012. PMID: 22972387 Free PMC article.
-
Extraction of bacterial RNA from soil: challenges and solutions.Microbes Environ. 2012;27(2):111-21. doi: 10.1264/jsme2.me11304. Microbes Environ. 2012. PMID: 22791042 Free PMC article. Review.
-
Functional Prediction of Biological Profile During Eutrophication in Marine Environment.Bioinform Biol Insights. 2022 Jan 5;16:11779322211063993. doi: 10.1177/11779322211063993. eCollection 2022. Bioinform Biol Insights. 2022. PMID: 35023908 Free PMC article.
-
Molecular techniques in the biotechnological fight against halogenated compounds in anoxic environments.Microb Biotechnol. 2012 May;5(3):347-67. doi: 10.1111/j.1751-7915.2011.00313.x. Epub 2011 Nov 9. Microb Biotechnol. 2012. PMID: 22070763 Free PMC article. Review.
References
-
- Allen, D. E., G. Kingston, H. Rennenberg, R. C. Dalal, and S. Schmidt. 2010. Effect of nitrogen fertilizer management and waterlogging on nitrous oxide emission from subtropical sugarcane soils. Agric. Ecosyst. Environ. 136:209-217.
-
- Campbell, E. J., P. M. Schenk, K. Kazan, I. A. Penninckx, J. P. Anderson, D. J. Maclean, B. P. Cammue, P. R. Ebert, and J. M. Manners. 2003. Pathogen-responsive expression of a putative ATP-binding cassette transporter gene conferring resistance to the diterpenoid sclareol is regulated by multiple defense signaling pathways in Arabidopsis. Plant Physiol. 133:1272-1284. - PMC - PubMed
-
- Chapman, S., P. M. Schenk, K. Kazan, and J. M. Manners. 2002. Using biplots to interpret gene expression patterns in plants. Bioinformatics 18:202-204. - PubMed
-
- Dalal, R. C., W. Wang, G. P. Robertson, and W. J. Parton. 2003. Nitrous oxide emission from Australian agricultural lands and mitigation options: a review. Aust. J. Soil Res. 41:165-196.
-
- Dalal, R. C., and D. E. Allen. 2008. Greenhouse gas fluxes from natural ecosystems (Turner review no. 18). Aust. J. Bot. 56:369-407.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

