Prediction of the clinical outcome in invasive candidiasis patients based on molecular fingerprints of five anti-Candida antibodies in serum
- PMID: 20860995
- PMCID: PMC3013459
- DOI: 10.1074/mcp.M110.004010
Prediction of the clinical outcome in invasive candidiasis patients based on molecular fingerprints of five anti-Candida antibodies in serum
Abstract
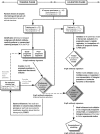
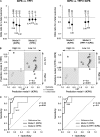
Better prognostic predictors for invasive candidiasis (IC) are needed to tailor and individualize therapeutic decision-making and minimize its high morbidity and mortality. We investigated whether molecular profiling of IgG-antibody response to the whole soluble Candida proteome could reveal a prognostic signature that may serve to devise a clinical-outcome prediction model for IC and contribute to known IC prognostic factors. By serological proteome analysis and data-mining procedures, serum 31-IgG antibody-reactivity patterns were examined in 45 IC patients randomly split into training and test sets. Within the training cohort, unsupervised two-way hierarchical clustering and principal-component analyses segregated IC patients into two antibody-reactivity subgroups with distinct prognoses that were unbiased by traditional IC prognostic factors and other patients-related variables. Supervised discriminant analysis with leave-one-out cross-validation identified a five-IgG antibody-reactivity signature as the most simplified and accurate IC clinical-outcome predictor, from which an IC prognosis score (ICPS) was derived. Its robustness was confirmed in the test set. Multivariate logistic-regression and receiver-operating-characteristic curve analyses demonstrated that the ICPS was able to accurately discriminate IC patients at high risk for death from those at low risk and outperformed conventional IC prognostic factors. Further validation of the five-IgG antibody-reactivity signature on a multiplexed immunoassay supported the serological proteome analysis results. The five IgG antibodies incorporated in the ICPS made biologic sense and were associated either with good-prognosis and protective patterns (those to Met6p, Hsp90p, and Pgk1p, putative Candida virulence factors and antiapoptotic mediators) or with poor-prognosis and risk patterns (those to Ssb1p and Gap1p/Tdh3p, potential Candida proapoptotic mediators). We conclude that the ICPS, with additional refinement in future larger prospective cohorts, could be applicable to reliably predict patient clinical-outcome for individualized therapy of IC. Our data further provide insights into molecular mechanisms that may influence clinical outcome in IC and uncover potential targets for vaccine design and immunotherapy against IC.
Figures







Similar articles
-
Serum antibody signature directed against Candida albicans Hsp90 and enolase detects invasive candidiasis in non-neutropenic patients.J Proteome Res. 2014 Nov 7;13(11):5165-84. doi: 10.1021/pr500681x. Epub 2014 Oct 14. J Proteome Res. 2014. PMID: 25377742
-
Decoding serological response to Candida cell wall immunome into novel diagnostic, prognostic, and therapeutic candidates for systemic candidiasis by proteomic and bioinformatic analyses.Mol Cell Proteomics. 2006 Jan;5(1):79-96. doi: 10.1074/mcp.M500243-MCP200. Epub 2005 Sep 28. Mol Cell Proteomics. 2006. PMID: 16195222
-
IgG1 anti-cell wall and IgG2 anti-phosphopeptidomannan antibodies in the diagnosis of invasive candidiasis and heavy Candida colonization.Med Mycol. 2015 Sep;53(7):725-35. doi: 10.1093/mmy/myv037. Epub 2015 Jul 10. Med Mycol. 2015. PMID: 26162476
-
Diagnostic accuracy of Candida albicans germ tube antibody for invasive candidiasis: systematic review and meta-analysis.Diagn Microbiol Infect Dis. 2019 Apr;93(4):339-345. doi: 10.1016/j.diagmicrobio.2018.10.017. Epub 2018 Nov 6. Diagn Microbiol Infect Dis. 2019. PMID: 30552034
-
Diagnosing Invasive Candidiasis.J Clin Microbiol. 2018 Apr 25;56(5):e01909-17. doi: 10.1128/JCM.01909-17. Print 2018 May. J Clin Microbiol. 2018. PMID: 29444828 Free PMC article. Review.
Cited by
-
Mass Spectrometry-Based Proteomic and Immunoproteomic Analyses of the Candida albicans Hyphal Secretome Reveal Diagnostic Biomarker Candidates for Invasive Candidiasis.J Fungi (Basel). 2021 Jun 23;7(7):501. doi: 10.3390/jof7070501. J Fungi (Basel). 2021. PMID: 34201883 Free PMC article.
-
Candida albicans Shaving to Profile Human Serum Proteins on Hyphal Surface.Front Microbiol. 2015 Dec 8;6:1343. doi: 10.3389/fmicb.2015.01343. eCollection 2015. Front Microbiol. 2015. PMID: 26696967 Free PMC article.
-
What Room for Two-Dimensional Gel-Based Proteomics in a Shotgun Proteomics World?Proteomes. 2020 Aug 6;8(3):17. doi: 10.3390/proteomes8030017. Proteomes. 2020. PMID: 32781532 Free PMC article. Review.
-
Top-down characterization data on the speciation of the Candida albicans immunome in candidemia.Data Brief. 2015 Dec 11;6:257-61. doi: 10.1016/j.dib.2015.11.054. eCollection 2016 Mar. Data Brief. 2015. PMID: 26862568 Free PMC article.
-
Moonlighting Proteins at the Candidal Cell Surface.Microorganisms. 2020 Jul 14;8(7):1046. doi: 10.3390/microorganisms8071046. Microorganisms. 2020. PMID: 32674422 Free PMC article. Review.
References
-
- Pappas P. G. (2006) Invasive candidiasis. Infect. Dis. Clin. North Am. 20, 485–506 - PubMed
-
- Eggimann P., Garbino J., Pittet D. (2003) Epidemiology of Candida species infections in critically ill non-immunosuppressed patients. Lancet Infect. Dis. 3, 685–702 - PubMed
-
- Falagas M. E., Apostolou K. E., Pappas V. D. (2006) Attributable mortality of candidemia: a systematic review of matched cohort and case-control studies. Eur. J. Clin. Microbiol. Infect. Dis. 25, 419–425 - PubMed
-
- Uzun O., Anaissie E. J. (2000) Predictors of outcome in cancer patients with candidemia. Ann. Oncol. 11, 1517–1521 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

