Evidence of functional protein dynamics from X-ray crystallographic ensembles
- PMID: 20865158
- PMCID: PMC2928775
- DOI: 10.1371/journal.pcbi.1000911
Evidence of functional protein dynamics from X-ray crystallographic ensembles
Abstract
It is widely recognized that representing a protein as a single static conformation is inadequate to describe the dynamics essential to the performance of its biological function. We contrast the amino acid displacements below and above the protein dynamical transition temperature, T(D)∼215K, of hen egg white lysozyme using X-ray crystallography ensembles that are analyzed by molecular dynamics simulations as a function of temperature. We show that measuring structural variations across an ensemble of X-ray derived models captures the activation of conformational states that are of functional importance just above T(D), and they remain virtually identical to structural motions measured at 300K. Our results highlight the ability to observe functional structural variations across an ensemble of X-ray crystallographic data, and that residue fluctuations measured in MD simulations at room temperature are in quantitative agreement with the experimental observable.
Conflict of interest statement
JZR was employed by the Public Library of Science at the time of submission.
Figures
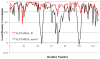



References
-
- Rasmussen BF, Stock AM, Ringe D, Petsko GA. Crystalline Ribonuclease-A Loses Function Below the Dynamic Transition at 220-K. Nature. 1992;357:423–424. - PubMed
-
- Bizzarri AR, Cannistraro S. Molecular Dynamics of Water at the Protein–Solvent Interface. J Phys Chem B. 2002;106:6617–6633.
-
- Lee AL, Wand AJ. Microscopic origins of entropy, heat capacity and the glass transition in proteins. Nature. 2001;411:501–504. - PubMed
-
- Born B, Kim SJ, Ebbinghaus S, Gruebele M, Havenith M. The terahertz dance of water with the proteins: the effect of protein flexibility on the dynamical hydration shell of ubiquitin. Faraday Discuss. 2009;141:161–173. - PubMed

