Burkholderia type VI secretion systems have distinct roles in eukaryotic and bacterial cell interactions
- PMID: 20865170
- PMCID: PMC2928800
- DOI: 10.1371/journal.ppat.1001068
Burkholderia type VI secretion systems have distinct roles in eukaryotic and bacterial cell interactions
Abstract
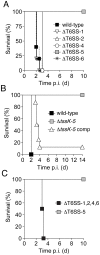
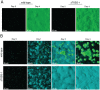
Bacteria that live in the environment have evolved pathways specialized to defend against eukaryotic organisms or other bacteria. In this manuscript, we systematically examined the role of the five type VI secretion systems (T6SSs) of Burkholderia thailandensis (B. thai) in eukaryotic and bacterial cell interactions. Consistent with phylogenetic analyses comparing the distribution of the B. thai T6SSs with well-characterized bacterial and eukaryotic cell-targeting T6SSs, we found that T6SS-5 plays a critical role in the virulence of the organism in a murine melioidosis model, while a strain lacking the other four T6SSs remained as virulent as the wild-type. The function of T6SS-5 appeared to be specialized to the host and not related to an in vivo growth defect, as ΔT6SS-5 was fully virulent in mice lacking MyD88. Next we probed the role of the five systems in interbacterial interactions. From a group of 31 diverse bacteria, we identified several organisms that competed less effectively against wild-type B. thai than a strain lacking T6SS-1 function. Inactivation of T6SS-1 renders B. thai greatly more susceptible to cell contact-induced stasis by Pseudomonas putida, Pseudomonas fluorescens and Serratia proteamaculans-leaving it 100- to 1000-fold less fit than the wild-type in competition experiments with these organisms. Flow cell biofilm assays showed that T6S-dependent interbacterial interactions are likely relevant in the environment. B. thai cells lacking T6SS-1 were rapidly displaced in mixed biofilms with P. putida, whereas wild-type cells persisted and overran the competitor. Our data show that T6SSs within a single organism can have distinct functions in eukaryotic versus bacterial cell interactions. These systems are likely to be a decisive factor in the survival of bacterial cells of one species in intimate association with those of another, such as in polymicrobial communities present both in the environment and in many infections.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





References
-
- Riley MA, Wertz JE. Bacteriocins: evolution, ecology, and application. Annu Rev Microbiol. 2002;56:117–137. - PubMed
-
- Cornelis GR. The type III secretion injectisome. Nat Rev Microbiol. 2006;4:811–825. - PubMed
-
- Bingle LE, Bailey CM, Pallen MJ. Type VI secretion: a beginner's guide. Curr Opin Microbiol. 2008;11:3–8. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

