A system for the analysis of BKV non-coding control regions: application to clinical isolates from an HIV/AIDS patient
- PMID: 20869740
- PMCID: PMC2957895
- DOI: 10.1016/j.virol.2010.08.032
A system for the analysis of BKV non-coding control regions: application to clinical isolates from an HIV/AIDS patient
Abstract
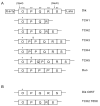
The human polyomavirus BK virus (BKV) is an important opportunistic pathogen whose disease prevalence continues to increase with the growing immunocompromised population. To date, the major determinant of replication in cell culture has not been formally proven. BKV exists as archetype virus and rearranged variants, which are classified based on the DNA sequence of their non-coding control regions (NCCRs). The archetype BKV NCCR is divided into five blocks of sequence and rearranged variants contain deletions and duplications of these blocks. In this study, a genetic system was developed and used to identify the major determinant of replication ability in primary renal proximal tubule epithelial cells, the natural host cell of BKV. This system was also used to analyze NCCR variants isolated from an immunocompromised patient which contain assorted rearrangement patterns and functional differences. This study solidifies the NCCR as the major genetic determinant of BKV replication ability in vitro.
Copyright © 2010 Elsevier Inc. All rights reserved.
Figures

References
-
- Chesters PM, Heritage J, McCance DJ. Persistence of DNA sequences of BK virus and JC virus in normal human tissues and in diseased tissues. J Infect Dis. 1983;147:676–84. - PubMed
-
- Cubukcu-Dimopulo O, Greco A, Kumar A, Karluk D, Mittal K, Jagirdar J. BK virus infection in AIDS. Am J Surg Pathol. 2000;24:145–9. - PubMed
-
- de Villiers J, Schaffner W, Tyndall C, Lupton S, Kamen R. Polyoma virus DNA replication requires an enhancer. Nature. 1984;312:242–6. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

