Control of the Staphylococcus aureus toxic shock tst promoter by the global regulator SarA
- PMID: 20870770
- PMCID: PMC2976458
- DOI: 10.1128/JB.00146-10
Control of the Staphylococcus aureus toxic shock tst promoter by the global regulator SarA
Abstract
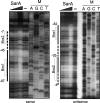
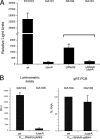
The Staphylococcus aureus SarA global regulator controls the expression of numerous virulence genes, often in conjunction with the agr quorum-sensing system and its effector RNA, RNAIII. In the present study, we have examined the role of both SarA and RNAIII on the regulation of the promoter of tst, encoding staphylococcal superantigen toxic shock syndrome toxin 1 (TSST-1). In vitro DNA-protein interaction studies with purified SarA using gel shift and DNase I protection assays revealed one strong SarA binding site and evidence for a weaker site nearby within the minimal 400-bp promoter region upstream of tst. In vivo analysis of tst promoter activation using a p(tst)-luxAB reporter inserted in the chromosome revealed partial but not complete loss of tst expression in a Δhld-RNAIII strain. In contrast, disruption of sarA abrogated tst expression. No significant tst expression was found for the double Δhld-RNAIII-ΔsarA mutant. Introduction of a plasmid containing cloned hld-RNAIII driven by a non-agr-dependent promoter, p(HU), into isogenic parental wild-type or ΔsarA strains showed comparable levels of RNAIII detected by quantitative reverse transcription-PCR (qRT-PCR) but a two-log(10) reduction in p(tst)-luxAB reporter expression in the ΔsarA strain, arguing that RNAIII levels alone are not strictly determinant for tst expression. Collectively, our results indicate that SarA binds directly to the tst promoter and that SarA plays a significant and direct role in the expression of tst.
Figures




References
-
- Blevins, J. S., A. F. Gillaspy, T. M. Rechtin, B. K. Hurlburt, and M. S. Smeltzer. 1999. The staphylococcal accessory regulator (sar) represses transcription of the Staphylococcus aureus collagen adhesin gene (cna) in an agr-independent manner. Mol. Microbiol. 33:317-326. - PubMed
-
- Chan, P. F., and S. J. Foster. 1998. The role of environmental factors in the regulation of virulence-determinant expression in Staphylococcus aureus 8325-4. Microbiology 144(Pt. 9):2469-2479. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

