Coarse-grained model for simulation of RNA three-dimensional structures
- PMID: 20883011
- PMCID: PMC2989335
- DOI: 10.1021/jp104926t
Coarse-grained model for simulation of RNA three-dimensional structures
Abstract
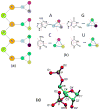
The accurate prediction of an RNA's three-dimensional structure from its "primary structure" will have a tremendous influence on the experimental design and its interpretation and ultimately our understanding of the many functions of RNA. This paper presents a general coarse-grained (CG) potential for modeling RNA 3-D structures. Each nucleotide is represented by five pseudo atoms, two for the backbone (one for the phosphate and another for the sugar) and three for the base to represent base-stacking interactions. The CG potential has been parametrized from statistical analysis of 688 RNA experimental structures. Molecular dynamic simulations of 15 RNA molecules with the length of 12-27 nucleotides have been performed using the CG potential, with performance comparable to that from all-atom simulations. For ~75% of systems tested, simulated annealing led to native-like structures at least once out of multiple repeated runs. Furthermore, with weak distance restraints based on the knowledge of three to five canonical Watson-Crick pairs, all 15 RNAs tested are successfully folded to within 6.5 Å of native structures using the CG potential and simulated annealing. The results reveal that with a limited secondary structure model the current CG potential can reliably predict the 3-D structures for small RNA molecules. We also explored an all-atom force field to construct atomic structures from the CG simulations.
Figures


Similar articles
-
RNA 3D structure prediction by using a coarse-grained model and experimental data.J Phys Chem B. 2013 Mar 21;117(11):3135-44. doi: 10.1021/jp400751w. Epub 2013 Mar 11. J Phys Chem B. 2013. PMID: 23438338
-
Modeling Noncanonical RNA Base Pairs by a Coarse-Grained IsRNA2 Model.J Phys Chem B. 2021 Nov 4;125(43):11907-11915. doi: 10.1021/acs.jpcb.1c07288. Epub 2021 Oct 25. J Phys Chem B. 2021. PMID: 34694128 Free PMC article.
-
Molecular dynamics and quantum mechanics of RNA: conformational and chemical change we can believe in.Acc Chem Res. 2010 Jan 19;43(1):40-7. doi: 10.1021/ar900093g. Acc Chem Res. 2010. PMID: 19754142 Free PMC article.
-
RNA 3D Structure Prediction Using Coarse-Grained Models.Front Mol Biosci. 2021 Jul 2;8:720937. doi: 10.3389/fmolb.2021.720937. eCollection 2021. Front Mol Biosci. 2021. PMID: 34277713 Free PMC article. Review.
-
RNA structure and dynamics: a base pairing perspective.Prog Biophys Mol Biol. 2013 Nov;113(2):264-83. doi: 10.1016/j.pbiomolbio.2013.07.003. Epub 2013 Jul 23. Prog Biophys Mol Biol. 2013. PMID: 23891726 Review.
Cited by
-
Prediction of solution properties and dynamics of RNAs by means of Brownian dynamics simulation of coarse-grained models: Ribosomal 5S RNA and phenylalanine transfer RNA.BMC Biophys. 2015 Dec 1;8:11. doi: 10.1186/s13628-015-0025-7. eCollection 2015. BMC Biophys. 2015. PMID: 26629336 Free PMC article.
-
Gay-Berne and electrostatic multipole based coarse-grain potential in implicit solvent.J Chem Phys. 2011 Oct 21;135(15):155104. doi: 10.1063/1.3651626. J Chem Phys. 2011. PMID: 22029338 Free PMC article.
-
Structure Prediction: New Insights into Decrypting Long Noncoding RNAs.Int J Mol Sci. 2016 Jan 21;17(1):132. doi: 10.3390/ijms17010132. Int J Mol Sci. 2016. PMID: 26805815 Free PMC article. Review.
-
Computational approaches to RNA structure prediction, analysis, and design.Curr Opin Struct Biol. 2011 Jun;21(3):306-18. doi: 10.1016/j.sbi.2011.03.015. Epub 2011 Apr 21. Curr Opin Struct Biol. 2011. PMID: 21514143 Free PMC article. Review.
-
Topological constraints of RNA pseudoknotted and loop-kissing motifs: applications to three-dimensional structure prediction.Nucleic Acids Res. 2020 Jul 9;48(12):6503-6512. doi: 10.1093/nar/gkaa463. Nucleic Acids Res. 2020. PMID: 32491164 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

