The L2a element is a mouse CD8 silencer that interacts with MAR-binding proteins SATB1 and CDP
- PMID: 20884053
- PMCID: PMC2996921
- DOI: 10.1016/j.molimm.2010.08.014
The L2a element is a mouse CD8 silencer that interacts with MAR-binding proteins SATB1 and CDP
Abstract
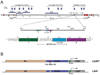
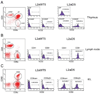
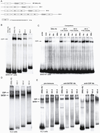
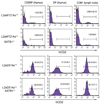
Previous transgenic-reporter and targeted-deletion studies indicate that the subset-specific expression of CD8αβ heterodimers is controlled by multiple enhancer activities, since no silencer elements had been found within the locus. We have identified such a silencer as L2a, a previously characterized ∼ 220 bp nuclear matrix associating region (MAR) located ∼ 4.5 kb upstream of CD8α. L2a transgenes driven by the E8(I) enhancer showed no reporter expression in thymic subsets or T cells in splenic, inguinal and mesenteric lymph node peripheral T cells. Deletion of L2a resulted in significant reporter de-repression, even in the CD4(+)CD8(+) double positive (DP) thymocyte population. L2a contains binding sites for two MAR-interacting proteins, SATB1 and CDP. We found that that binding of these factors was markedly influenced by the content and spacing of L2a sub-motifs (L and S) and that SATB1 binds preferentially to the L motif both in vitro and in vivo. A small fraction of the transgenic CD8 single positive (SP) thymocytes and peripheral CD8(+) T cells bypassed L2a-silencing to give rise to variegated expression of the transgenic reporter. Crossing the L2a-containing transgene onto a SATB1 knockdown background enhanced variegated expression, suggesting that SATB1 is critical in overcoming L2a-silenced transcription.
Copyright © 2010 Elsevier Ltd. All rights reserved.
Conflict of interest statement
All authors concur with the submission and that the material submitted for publication has not been previously reported and is not under consideration for publication elsewhere; the authors have no financial conflict of interest.
Figures

Similar articles
-
SATB1-mediated chromatin landscape in T cells.Nucleus. 2020 Dec;11(1):117-131. doi: 10.1080/19491034.2020.1775037. Nucleus. 2020. PMID: 33356851 Free PMC article. Review.
-
Interaction of the nuclear matrix-associated region (MAR)-binding proteins, SATB1 and CDP/Cux, with a MAR element (L2a) in an upstream regulatory region of the mouse CD8a gene.J Biol Chem. 1997 Jul 18;272(29):18440-52. doi: 10.1074/jbc.272.29.18440. J Biol Chem. 1997. PMID: 9218488
-
A role for SATB1, a nuclear matrix association region-binding protein, in the development of CD8SP thymocytes and peripheral T lymphocytes.J Immunol. 2005 Apr 15;174(8):4745-52. doi: 10.4049/jimmunol.174.8.4745. J Immunol. 2005. PMID: 15814699
-
A nuclear matrix attachment region upstream of the T cell receptor beta gene enhancer binds Cux/CDP and SATB1 and modulates enhancer-dependent reporter gene expression but not endogenous gene expression.J Biol Chem. 1998 Nov 6;273(45):29838-46. doi: 10.1074/jbc.273.45.29838. J Biol Chem. 1998. PMID: 9792700
-
A mini review of MAR-binding proteins.Mol Biol Rep. 2010 Oct;37(7):3553-60. doi: 10.1007/s11033-010-0003-8. Epub 2010 Feb 22. Mol Biol Rep. 2010. PMID: 20174991 Review.
Cited by
-
Short tandem repeats are important contributors to silencer elements in T cells.Nucleic Acids Res. 2023 Jun 9;51(10):4845-4866. doi: 10.1093/nar/gkad187. Nucleic Acids Res. 2023. PMID: 36929452 Free PMC article.
-
SATB1-mediated chromatin landscape in T cells.Nucleus. 2020 Dec;11(1):117-131. doi: 10.1080/19491034.2020.1775037. Nucleus. 2020. PMID: 33356851 Free PMC article. Review.
-
Transcriptional control of CD4 and CD8 coreceptor expression during T cell development.Cell Mol Life Sci. 2013 Dec;70(23):4537-53. doi: 10.1007/s00018-013-1393-2. Epub 2013 Jun 21. Cell Mol Life Sci. 2013. PMID: 23793512 Free PMC article. Review.
-
An integrative computational systems biology approach identifies differentially regulated dynamic transcriptome signatures which drive the initiation of human T helper cell differentiation.BMC Genomics. 2012 Oct 30;13:572. doi: 10.1186/1471-2164-13-572. BMC Genomics. 2012. PMID: 23110343 Free PMC article.
-
Heritable Gene Regulation in the CD4:CD8 T Cell Lineage Choice.Front Immunol. 2017 Mar 22;8:291. doi: 10.3389/fimmu.2017.00291. eCollection 2017. Front Immunol. 2017. PMID: 28382035 Free PMC article. Review.
References
-
- Banan M, Rojas IC, Lee WH, King HL, Harriss JV, Kobayashi R, Webb CF, Gottlieb PD. Interaction of the nuclear matrix-associated region (MAR)-binding proteins, SATB1 and CDP/Cux, with a MAR element (L2a) in an upstream regulatory region of the mouse CD8a gene. J Biol Chem. 1997;272:18440–18452. - PubMed
-
- Blasquez VC, Sperry AO, Cockerill PN, Garrard WT. Protein:DNA interactions at chromosomal loop attachment sites. Genome. 1989;31:503–509. - PubMed
-
- Cai S, Han HJ, Kohwi-Shigematsu T. Tissue-specific nuclear architecture and gene expression regulated by SATB1. Nat Genet. 2003;34:42–51. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

