Implementing a rational and consistent nomenclature for serine/arginine-rich protein splicing factors (SR proteins) in plants
- PMID: 20884799
- PMCID: PMC2965536
- DOI: 10.1105/tpc.110.078352
Implementing a rational and consistent nomenclature for serine/arginine-rich protein splicing factors (SR proteins) in plants
Abstract
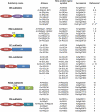
Growing interest in alternative splicing in plants and the extensive sequencing of new plant genomes necessitate more precise definition and classification of genes coding for splicing factors. SR proteins are a family of RNA binding proteins, which function as essential factors for constitutive and alternative splicing. We propose a unified nomenclature for plant SR proteins, taking into account the newly revised nomenclature of the mammalian SR proteins and a number of plant-specific properties of the plant proteins. We identify six subfamilies of SR proteins in Arabidopsis thaliana and rice (Oryza sativa), three of which are plant specific. The proposed subdivision of plant SR proteins into different subfamilies will allow grouping of paralogous proteins and simple assignment of newly discovered SR orthologs from other plant species and will promote functional comparisons in diverse plant species.
Figures
Similar articles
-
Comparative analysis of serine/arginine-rich proteins across 27 eukaryotes: insights into sub-family classification and extent of alternative splicing.PLoS One. 2011;6(9):e24542. doi: 10.1371/journal.pone.0024542. Epub 2011 Sep 14. PLoS One. 2011. PMID: 21935421 Free PMC article.
-
Multiplex CRISPR Mutagenesis of the Serine/Arginine-Rich (SR) Gene Family in Rice.Genes (Basel). 2019 Aug 7;10(8):596. doi: 10.3390/genes10080596. Genes (Basel). 2019. PMID: 31394891 Free PMC article.
-
The serine/arginine-rich protein family in rice plays important roles in constitutive and alternative splicing of pre-mRNA.Plant Cell. 2006 Jan;18(1):146-58. doi: 10.1105/tpc.105.037069. Epub 2005 Dec 9. Plant Cell. 2006. PMID: 16339852 Free PMC article.
-
Plant serine/arginine-rich proteins: roles in precursor messenger RNA splicing, plant development, and stress responses.Wiley Interdiscip Rev RNA. 2011 Nov-Dec;2(6):875-89. doi: 10.1002/wrna.98. Epub 2011 Jul 15. Wiley Interdiscip Rev RNA. 2011. PMID: 21766458 Review.
-
A role for SR proteins in plant stress responses.Plant Signal Behav. 2011 Jan;6(1):49-54. doi: 10.4161/psb.6.1.14063. Epub 2011 Jan 1. Plant Signal Behav. 2011. PMID: 21258207 Free PMC article. Review.
Cited by
-
Changes in Alternative Splicing in Response to Domestication and Polyploidization in Wheat.Plant Physiol. 2020 Dec;184(4):1955-1968. doi: 10.1104/pp.20.00773. Epub 2020 Oct 13. Plant Physiol. 2020. PMID: 33051269 Free PMC article.
-
The spliceosome-activating complex: molecular mechanisms underlying the function of a pleiotropic regulator.Front Plant Sci. 2012 Jan 26;3:9. doi: 10.3389/fpls.2012.00009. eCollection 2012. Front Plant Sci. 2012. PMID: 22639636 Free PMC article.
-
SRSF3 Expression Serves as a Potential Biomarker for Prognostic and Immune Response in Pan-Cancer.Front Oncol. 2022 Apr 14;12:808530. doi: 10.3389/fonc.2022.808530. eCollection 2022. Front Oncol. 2022. PMID: 35494088 Free PMC article.
-
Transportin-SR is required for proper splicing of resistance genes and plant immunity.PLoS Genet. 2011 Jun;7(6):e1002159. doi: 10.1371/journal.pgen.1002159. Epub 2011 Jun 30. PLoS Genet. 2011. PMID: 21738492 Free PMC article.
-
Plant serine/arginine-rich proteins: versatile players in RNA processing.Planta. 2023 May 5;257(6):109. doi: 10.1007/s00425-023-04132-0. Planta. 2023. PMID: 37145304 Review.
References
-
- Golovkin M., Reddy A.S. (1999). An SC35-like protein and a novel serine/arginine-rich protein interact with Arabidopsis U1-70K protein. J. Biol. Chem. 274: 36428–36438 - PubMed
-
- Huang Y., Steitz J.A. (2005). SRprises along a messenger's journey. Mol. Cell 17: 613–615 - PubMed
-
- Iida K., Go M. (2006). Survey of conserved alternative splicing events of mRNAs encoding SR proteins in land plants. Mol. Biol. Evol. 23: 1085–1094 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

