Activation-induced cytidine deaminase targets DNA at sites of RNA polymerase II stalling by interaction with Spt5
- PMID: 20887897
- PMCID: PMC2993080
- DOI: 10.1016/j.cell.2010.09.017
Activation-induced cytidine deaminase targets DNA at sites of RNA polymerase II stalling by interaction with Spt5
Abstract
Activation-induced cytidine deaminase (AID) initiates antibody gene diversification by creating U:G mismatches. However, AID is not specific for antibody genes; Off-target lesions can activate oncogenes or cause chromosome translocations. Despite its importance in these transactions little is known about how AID finds its targets. We performed an shRNA screen to identify factors required for class switch recombination (CSR) of antibody loci. We found that Spt5, a factor associated with stalled RNA polymerase II (Pol II) and single stranded DNA (ssDNA), is required for CSR. Spt5 interacts with AID, it facilitates association between AID and Pol II, and AID recruitment to its Ig and non-Ig targets. ChIP-seq experiments reveal that Spt5 colocalizes with AID and stalled Pol II. Further, Spt5 accumulation at sites of Pol II stalling is predictive of AID-induced mutation. We propose that AID is targeted to sites of Pol II stalling in part via its association with Spt5.
Copyright © 2010 Elsevier Inc. All rights reserved.
Figures



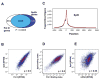


References
-
- Amir-Zilberstein L, Dikstein R. Interplay between E-box and NF-kappaB in regulation of A20 gene by DRB sensitivity-inducing factor (DSIF) J Biol Chem. 2008;283:1317–1323. - PubMed
-
- Andrulis ED, Werner J, Nazarian A, Erdjument-Bromage H, Tempst P, Lis JT. The RNA processing exosome is linked to elongating RNA polymerase II in Drosophila. Nature. 2002;420:837–841. - PubMed
-
- Basu U, Chaudhuri J, Alpert C, Dutt S, Ranganath S, Li G, Schrum JP, Manis JP, Alt FW. The AID antibody diversification enzyme is regulated by protein kinase A phosphorylation. Nature. 2005;438:508–511. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

