Recruitment of genes and enzymes conferring resistance to the nonnatural toxin bromoacetate
- PMID: 20921376
- PMCID: PMC2964223
- DOI: 10.1073/pnas.1007559107
Recruitment of genes and enzymes conferring resistance to the nonnatural toxin bromoacetate
Abstract
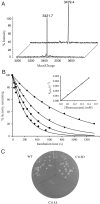
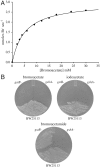
Microbial niches contain toxic chemicals capable of forcing organisms into periods of intense natural selection to afford survival. Elucidating the mechanisms by which microbes evade environmental threats has direct relevance for understanding and combating the rise of antibiotic resistance. In this study we used a toxic small-molecule, bromoacetate, to model the selective pressures imposed by antibiotics and anthropogenic toxins. We report the results of genetic selection experiments that identify nine genes from Escherichia coli whose overexpression affords survival in the presence of a normally lethal concentration of bromoacetate. Eight of these genes encode putative transporters or transmembrane proteins, while one encodes the essential peptidoglycan biosynthetic enzyme, UDP-N-acetylglucosamine enolpyruvoyl transferase (MurA). Biochemical studies demonstrate that the primary physiological target of bromoacetate is MurA, which becomes irreversibly inactivated via alkylation of a critical active-site cysteine. We also screened a comprehensive library of E. coli single-gene deletion mutants and identified 63 strains displaying increased susceptibility to bromoacetate. One hypersensitive bacterium lacks yliJ, a gene encoding a predicted glutathione transferase. Herein, YliJ is shown to catalyze the glutathione-dependent dehalogenation of bromoacetate with a k(cat)/K(m) value of 5.4 × 10(3) M(-1) s(-1). YliJ displays exceptional substrate specificity and produces a rate enhancement exceeding 5 orders of magnitude, remarkable characteristics for reactivity with a nonnatural molecule. This study illustrates the wealth of intrinsic survival mechanisms that can be exploited by bacteria when they are challenged with toxins.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
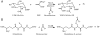
Similar articles
-
In vitro and in vivo functional activity of Chlamydia MurA, a UDP-N-acetylglucosamine enolpyruvyl transferase involved in peptidoglycan synthesis and fosfomycin resistance.J Bacteriol. 2003 Feb;185(4):1218-28. doi: 10.1128/JB.185.4.1218-1228.2003. J Bacteriol. 2003. PMID: 12562791 Free PMC article.
-
Boswellic acids, as novel inhibitor targeting peptidoglycan biosynthetic enzyme UDP-N-acetylglucosamine enolpyruvyl transferase (MurA) in Escherichia coli.Arch Microbiol. 2022 Jul 12;204(8):472. doi: 10.1007/s00203-022-03066-7. Arch Microbiol. 2022. PMID: 35819545
-
Identification of a novel UDP-N-acetylglucosamine enolpyruvyl transferase (MurA) from Vibrio fischeri that confers high fosfomycin resistance in Escherichia coli.Arch Microbiol. 2009 May;191(5):425-9. doi: 10.1007/s00203-009-0468-9. Epub 2009 Mar 11. Arch Microbiol. 2009. PMID: 19277606 Free PMC article.
-
Potential Inhibitors Targeting Escherichia coli UDP-N-Acetylglucosamine Enolpyruvyl Transferase (MurA): An Overview.Indian J Microbiol. 2022 Mar;62(1):11-22. doi: 10.1007/s12088-021-00988-6. Epub 2021 Oct 29. Indian J Microbiol. 2022. PMID: 35068599 Free PMC article. Review.
-
Advances in UDP-N-Acetylglucosamine Enolpyruvyl Transferase (MurA) Covalent Inhibition.Front Mol Biosci. 2022 Jul 20;9:889825. doi: 10.3389/fmolb.2022.889825. eCollection 2022. Front Mol Biosci. 2022. PMID: 35936791 Free PMC article. Review.
Cited by
-
Metabolic engineering strategies for L-Homoserine production in Escherichia coli.Microb Cell Fact. 2024 Dec 19;23(1):338. doi: 10.1186/s12934-024-02623-7. Microb Cell Fact. 2024. PMID: 39702271 Free PMC article. Review.
-
An alternate pathway of arsenate resistance in E. coli mediated by the glutathione S-transferase GstB.ACS Chem Biol. 2015 Mar 20;10(3):875-82. doi: 10.1021/cb500755j. Epub 2015 Jan 7. ACS Chem Biol. 2015. PMID: 25517993 Free PMC article.
-
Evolution of pollutant biodegradation.Appl Microbiol Biotechnol. 2025 Feb 4;109(1):36. doi: 10.1007/s00253-025-13418-0. Appl Microbiol Biotechnol. 2025. PMID: 39903283 Free PMC article. Review.
-
Structural and biochemical characterization of a glutathione transferase from the citrus canker pathogen Xanthomonas.Acta Crystallogr D Struct Biol. 2020 Aug 1;76(Pt 8):778-789. doi: 10.1107/S2059798320009274. Epub 2020 Jul 27. Acta Crystallogr D Struct Biol. 2020. PMID: 32744260 Free PMC article.
-
Substrate ambiguous enzymes within the Escherichia coli proteome offer different evolutionary solutions to the same problem.Mol Biol Evol. 2013 Sep;30(9):2001-12. doi: 10.1093/molbev/mst105. Epub 2013 May 31. Mol Biol Evol. 2013. PMID: 23728795 Free PMC article.
References
-
- Walsh CT. Molecular mechanisms that confer antibacterial drug resistance. Nature. 2000;406:775–781. - PubMed
-
- Davies J. Inactivation of antibiotics and the dissemination of resistance genes. Science. 1994;264:375–382. - PubMed
-
- Davies JE. Origins, acquisition and dissemination of antibiotic resistance determinants. Ciba Found Symp. 1997;207:15–27. - PubMed
-
- Hall BG. Predicting the evolution of antibiotic resistance genes. Nat Rev Microbiol. 2004;2:430–435. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

