Extreme genetic divergence is required for coreceptor switching in HIV-1 subtype C
- PMID: 20921899
- PMCID: PMC3006070
- DOI: 10.1097/QAI.0b013e3181f63906
Extreme genetic divergence is required for coreceptor switching in HIV-1 subtype C
Abstract
Background: Coreceptor switching from CCR5 to CXCR4 is less common in subtype C HIV-1 infection than in subtype B for reasons that are unclear. We have examined sequential virus samples from a subtype C-infected child who had evidence of coreceptor switching.
Methods: To examine HIV-1 envelope evolution towards CXCR4 usage, env sequences were correlated with phenotypic characteristics determined by entry assays, as well as the ability to use alternative coreceptors such as FPRL1, CCR3, CCR8 and others. The value of a phenotype predictor based on V3 sequences was also assessed.
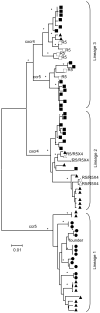
Results: Ninety-three sequences revealed 3 distinct coexistent virus lineages and only some members of one lineage evolved to use CXCR4. These lineages also had diverse alternative coreceptor patterns including the ability to use FPRL1, CCR3, CCR8, APJ, CMKLR1, RDC-1, CXCR6, CCR1, GPCR1, GPR15 and CCR6. Coreceptor switching was associated with extensive and rapid sequence divergence in the V1/V2 region in addition to V3 changes. Furthermore, interlineage recombination within the C2 region resulted in low predictability of a V3 sequence-based phenotype algorithm, and highlighted the importance of V1/V2 and V3 sequences in coreceptor usage.
Conclusion: These results suggest that the evolution to coreceptor switching in subtype C infection requires more mutations than other subtypes, and this contributes to the reduced incidence of R5X4 viruses.
Figures

References
-
- Richman DD, Bozzette SA. The impact of the syncytium-inducing phenotype of human immunodeficiency virus on disease progression. J Infect Dis. 1994;169(5):968–974. - PubMed
-
- Scarlatti G, Tresoldi E, Bjorndal A, et al. In vivo evolution of HIV-1 co-receptor usage and sensitivity to chemokine-mediated suppression. Nat Med. 1997;3(11):1259–1265. - PubMed
-
- Abebe A, Demissie D, Goudsmit J, et al. HIV-1 subtype C syncytium- and non-syncytium-inducing phenotypes and coreceptor usage among Ethiopian patients with AIDS. Aids. 1999 Jul 30;13(11):1305–1311. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

