Tubulin depolymerization may be an ancient biological motor
- PMID: 20930138
- PMCID: PMC2959111
- DOI: 10.1242/jcs.067611
Tubulin depolymerization may be an ancient biological motor
Abstract
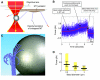
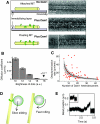
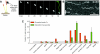
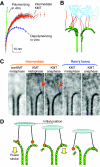
The motions of mitotic chromosomes are complex and show considerable variety across species. A wealth of evidence supports the idea that microtubule-dependent motor enzymes contribute to this variation and are important both for spindle formation and for the accurate completion of chromosome segregation. Motors that walk towards the spindle pole are, however, dispensable for at least some poleward movements of chromosomes in yeasts, suggesting that depolymerizing spindle microtubules can generate mitotic forces in vivo. Tubulin protofilaments that flare outward in association with microtubule shortening may be the origin of such forces, because they can move objects that are appropriately attached to a microtubule wall. For example, some kinetochore-associated proteins can couple experimental objects, such as microspheres, to shortening microtubules in vitro, moving them over many micrometers. Here, we review recent evidence about such phenomena, highlighting the force-generation mechanisms and different coupling strategies. We also consider bending filaments of the tubulin-like protein FtsZ, which form rings girding bacteria at their sites of cytokinesis. Mechanical similarities between these force-generation systems suggest a deep phylogenetic relationship between tubulin depolymerization in eukaryotic mitosis and FtsZ-mediated ring contraction in bacteria.
Figures


References
-
- Brinkley B. R., Stubblefield E. (1966). The fine structure of the kinetochore of a mammalian cell in vitro. Chromosoma 19, 28-43 - PubMed
-
- Cassimeris L., Inoue S., Salmon E. D. (1988). Microtubule dynamics in the chromosomal spindle fiber: analysis by fluorescence and high-resolution polarization microscopy. Cell Motil. Cytoskeleton 10, 185-196 - PubMed
-
- Cheeseman I. M., Chappie J. S., Wilson-Kubalek E. M., Desai A. (2006). The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell 127, 983-997 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

