Molecular detection of targeted major histocompatibility complex I-bound peptides using a probabilistic measure and nanospray MS3 on a hybrid quadrupole-linear ion trap
- PMID: 20932029
- PMCID: PMC2966287
- DOI: 10.1021/ac102387t
Molecular detection of targeted major histocompatibility complex I-bound peptides using a probabilistic measure and nanospray MS3 on a hybrid quadrupole-linear ion trap
Abstract
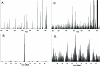
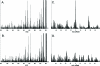
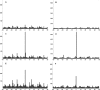
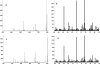
A nanospray MS(3) method deployed on a quadrupole linear ion trap hybrid can detect targeted peptides with high dynamic range and high sensitivity from complex mixtures without separations. The method uses a recognition algorithm that is a modification of the relative (Kullback-Leibler, KL) entropy characterization of probabilistic distance to detect if reference MS(3) fragmentation patterns are components of acquired MS(3) spectra. The recognition reflects the probabilistic structure of physical MS measurements unlike the Euclidean or inner product metrics widely used for comparing spectra. It capably handles spectra with a significant chemical ion background in contrast to the Euclidean metric or the direct relative entropy. The full nanospray MS(3) method allows both the detection and quantitation of targets without the need to obtain isotopically labeled standards. By avoiding chromatographic separations and its associated surface losses, the detection can be applied to complex samples on a very limited material scale. The methodology is illustrated by applications to the medically important problem of detecting targeted major histocompatibility complex (MHC) I associated peptides extracted from limited cell numbers.
Figures

Similar articles
-
Accurate quantitation of MHC-bound peptides by application of isotopically labeled peptide MHC complexes.J Proteomics. 2014 Sep 23;109:240-4. doi: 10.1016/j.jprot.2014.07.009. Epub 2014 Jul 19. J Proteomics. 2014. PMID: 25050860
-
High-performance liquid chromatography-tandem mass spectrometry with a new quadrupole/linear ion trap instrument.J Chromatogr A. 2003 Dec 5;1020(1):3-9. doi: 10.1016/s0021-9673(03)00426-6. J Chromatogr A. 2003. PMID: 14661752
-
Quadrupole ion trap mass spectrometry.Methods Enzymol. 1996;270:552-86. doi: 10.1016/s0076-6879(96)70025-3. Methods Enzymol. 1996. PMID: 8803984
-
Comparison of a preliminary procedure for the general unknown screening of drugs and toxic compounds using a quadrupole-linear ion-trap mass spectrometer with a liquid chromatography-mass spectrometry reference technique.J Chromatogr B Analyt Technol Biomed Life Sci. 2003 Jun 5;789(1):9-18. doi: 10.1016/s1570-0232(03)00071-0. J Chromatogr B Analyt Technol Biomed Life Sci. 2003. PMID: 12726839
-
Quantifying epitope presentation using mass spectrometry.Mol Immunol. 2015 Dec;68(2 Pt A):77-80. doi: 10.1016/j.molimm.2015.06.010. Epub 2015 Jun 25. Mol Immunol. 2015. PMID: 26118903 Review.
Cited by
-
Direct identification of an HPV-16 tumor antigen from cervical cancer biopsy specimens.Front Immunol. 2011 Dec 13;2:75. doi: 10.3389/fimmu.2011.00075. eCollection 2011. Front Immunol. 2011. PMID: 22566864 Free PMC article.
-
Computational resources for high-dimensional immune analysis from the Human Immunology Project Consortium.Nat Biotechnol. 2014 Feb;32(2):146-8. doi: 10.1038/nbt.2777. Epub 2014 Jan 19. Nat Biotechnol. 2014. PMID: 24441472 Free PMC article. No abstract available.
-
HLA-I immunopeptidome profiling of human cells infected with high-containment enveloped viruses.STAR Protoc. 2022 Dec 16;3(4):101910. doi: 10.1016/j.xpro.2022.101910. Epub 2022 Dec 8. STAR Protoc. 2022. PMID: 36595954 Free PMC article.
-
ALK peptide vaccination restores the immunogenicity of ALK-rearranged non-small cell lung cancer.Nat Cancer. 2023 Jul;4(7):1016-1035. doi: 10.1038/s43018-023-00591-2. Epub 2023 Jul 10. Nat Cancer. 2023. PMID: 37430060 Free PMC article.
-
TCR-mimic bispecific antibodies to target the HIV-1 reservoir.Proc Natl Acad Sci U S A. 2022 Apr 12;119(15):e2123406119. doi: 10.1073/pnas.2123406119. Epub 2022 Apr 8. Proc Natl Acad Sci U S A. 2022. PMID: 35394875 Free PMC article.
References
-
- Groothuis T.; Neefjes J. Curr. Top. Microbiol. Immunol. 2005, 300, 127–148. - PubMed
-
- Shastri N.; Cardinaud S.; Schwab S. R.; Serwold T.; Kunisawa J. Immunol. Rev. 2005, 207, 31–41. - PubMed
-
- Kloetzel P. M. Biochim. Biophys. Acta 2004, 1695, 225–233. - PubMed
-
- Lehner P. J.; Cresswell P. Curr. Opin. Immunol. 2004, 16, 82–89. - PubMed
-
- Yewdell J. W.; Reits E.; Neefjes J. Nat. Rev. Immunol. 2003, 3, 952–961. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

