Gene expression in a paleopolyploid: a transcriptome resource for the ciliate Paramecium tetraurelia
- PMID: 20932287
- PMCID: PMC3091696
- DOI: 10.1186/1471-2164-11-547
Gene expression in a paleopolyploid: a transcriptome resource for the ciliate Paramecium tetraurelia
Abstract
Background: The genome of Paramecium tetraurelia, a unicellular model that belongs to the ciliate phylum, has been shaped by at least 3 successive whole genome duplications (WGD). These dramatic events, which have also been documented in plants, animals and fungi, are resolved over evolutionary time by the loss of one duplicate for the majority of genes. Thanks to a low rate of large scale genome rearrangement in Paramecium, an unprecedented large number of gene duplicates of different ages have been identified, making this organism an outstanding model to investigate the evolutionary consequences of polyploidization. The most recent WGD, with 51% of pre-duplication genes still in 2 copies, provides a snapshot of a phase of rapid gene loss that is not accessible in more ancient polyploids such as yeast.
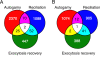
Results: We designed a custom oligonucleotide microarray platform for P. tetraurelia genome-wide expression profiling and used the platform to measure gene expression during 1) the sexual cycle of autogamy, 2) growth of new cilia in response to deciliation and 3) biogenesis of secretory granules after massive exocytosis. Genes that are differentially expressed during these time course experiments have expression patterns consistent with a very low rate of subfunctionalization (partition of ancestral functions between duplicated genes) in particular since the most recent polyploidization event.
Conclusions: A public transcriptome resource is now available for Paramecium tetraurelia. The resource has been integrated into the ParameciumDB model organism database, providing searchable access to the data. The microarray platform, freely available through NimbleGen Systems, provides a robust, cost-effective approach for genome-wide expression profiling in P. tetraurelia. The expression data support previous studies showing that at short evolutionary times after a whole genome duplication, gene dosage balance constraints and not functional change are the major determinants of gene retention.
Figures

References
-
- Sémon M, Wolfe KH. Consequences of genome duplication. Current Opinion in Genetics & Development. 2007;17:505–512. - PubMed
Publication types
MeSH terms
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

