Evolution and population structure of Salmonella enterica serovar Newport
- PMID: 20935094
- PMCID: PMC3008538
- DOI: 10.1128/JB.00969-10
Evolution and population structure of Salmonella enterica serovar Newport
Abstract
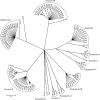
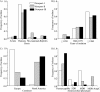
Salmonellosis caused by Salmonella enterica serovar Newport is a major global public health concern, particularly because S. Newport isolates that are resistant to multiple drugs (MDR), including third-generation cephalosporins (MDR-AmpC phenotype), have been commonly isolated from food animals. We analyzed 384 S. Newport isolates from various sources by a multilocus sequence typing (MLST) scheme to study the evolution and population structure of the serovar. These were compared to the population structure of S. enterica serovars Enteritidis, Kentucky, Paratyphi B, and Typhimurium. Our S. Newport collection fell into three lineages, Newport-I, Newport-II, and Newport-III, each of which contained multiple sequence types (STs). Newport-I has only a few STs, unlike Newport-II or Newport-III, and has possibly emerged recently. Newport-I is more prevalent among humans in Europe than in North America, whereas Newport-II is preferentially associated with animals. Two STs of Newport-II encompassed all MDR-AmpC isolates, suggesting recent global spread after the acquisition of the bla(CMY-2) gene. In contrast, most Newport-III isolates were from humans in North America and were pansusceptible to antibiotics. Newport was intermediate in population structure to the other serovars, which varied from a single monophyletic lineage in S. Enteritidis or S. Typhimurium to four discrete lineages within S. Paratyphi B. Both mutation and homologous recombination are responsible for diversification within each of these lineages, but the relative frequencies differed with the lineage. We conclude that serovars of S. enterica provide a variety of different population structures.
Figures


Similar articles
-
Comparison of multilocus sequence typing, pulsed-field gel electrophoresis, and antimicrobial susceptibility typing for characterization of Salmonella enterica serotype Newport isolates.J Clin Microbiol. 2006 Jul;44(7):2449-57. doi: 10.1128/JCM.00019-06. J Clin Microbiol. 2006. PMID: 16825363 Free PMC article.
-
Distribution of multidrug-resistant human isolates of MDR-ACSSuT Salmonella Typhimurium and MDR-AmpC Salmonella Newport in the United States, 2003-2005.Foodborne Pathog Dis. 2008 Oct;5(5):669-80. doi: 10.1089/fpd.2008.0111. Foodborne Pathog Dis. 2008. PMID: 18851677
-
Use of antibiotic susceptibility patterns and pulsed-field gel electrophoresis to compare historic and contemporary isolates of multi-drug-resistant Salmonella enterica subsp. enterica serovar Newport.Appl Environ Microbiol. 2004 Jan;70(1):318-23. doi: 10.1128/AEM.70.1.318-323.2004. Appl Environ Microbiol. 2004. PMID: 14711658 Free PMC article.
-
Population structure, origins and evolution of major Salmonella enterica clones.Infect Genet Evol. 2009 Sep;9(5):996-1005. doi: 10.1016/j.meegid.2009.04.011. Epub 2009 Apr 23. Infect Genet Evol. 2009. PMID: 19393770 Review.
-
Salmonella serovars from foodborne and waterborne diseases in Korea, 1998-2007: total isolates decreasing versus rare serovars emerging.J Korean Med Sci. 2010 Dec;25(12):1693-9. doi: 10.3346/jkms.2010.25.12.1693. Epub 2010 Nov 24. J Korean Med Sci. 2010. PMID: 21165281 Free PMC article. Review.
Cited by
-
Identification of Salmonella for public health surveillance using whole genome sequencing.PeerJ. 2016 Apr 5;4:e1752. doi: 10.7717/peerj.1752. eCollection 2016. PeerJ. 2016. PMID: 27069781 Free PMC article.
-
Salmonella enterica Phylogeny Based on Whole-Genome Sequencing Reveals Two New Clades and Novel Patterns of Horizontally Acquired Genetic Elements.mBio. 2018 Nov 27;9(6):e02303-18. doi: 10.1128/mBio.02303-18. mBio. 2018. PMID: 30482836 Free PMC article.
-
Worldwide Epidemiology of Salmonella Serovars in Animal-Based Foods: a Meta-analysis.Appl Environ Microbiol. 2019 Jul 1;85(14):e00591-19. doi: 10.1128/AEM.00591-19. Print 2019 Jul 15. Appl Environ Microbiol. 2019. PMID: 31053586 Free PMC article.
-
Multiple Food-Animal-Borne Route in Transmission of Antibiotic-Resistant Salmonella Newport to Humans.Front Microbiol. 2018 Jan 23;9:23. doi: 10.3389/fmicb.2018.00023. eCollection 2018. Front Microbiol. 2018. PMID: 29410657 Free PMC article.
-
Comparative genomic analysis reveals high intra-serovar plasticity within Salmonella Napoli isolated in 2005-2017.BMC Genomics. 2020 Mar 4;21(1):202. doi: 10.1186/s12864-020-6588-y. BMC Genomics. 2020. PMID: 32131727 Free PMC article.
References
-
- Alcaine, S. D., Y. Soyer, L. D. Warnick, W. L. Su, S. Sukhnananda, J. Richards, E. D. Fortes, P. McDonough, T. P. Root, N. B. Dumas, Y. Grohn, and M. Wiedmann. 2006. Multilocus sequence typing supports the hypothesis that cow- and human-associated Salmonella isolates represent distinct and overlapping populations. Appl. Environ. Microbiol. 72:7575-7585. - PMC - PubMed
-
- Alcaine, S. D., S. S. Sukhnanand, L. D. Warnick, W. L. Su, P. McGann, P. McDonough, and M. Wiedmann. 2005. Ceftiofur-resistant Salmonella strains isolated from dairy farms represent multiple widely distributed subtypes that evolved by independent horizontal gene transfer. Antimicrob. Agents Chemother. 49:4061-4067. - PMC - PubMed
-
- Arlet, G., T. J. Barrett, P. Butaye, A. Cloeckaert, M. R. Mulvey, and D. G. White. 2006. Salmonella resistant to extended-spectrum cephalosporins: prevalence and epidemiology. Microbes Infect. 8:1945-1954. - PubMed
-
- Barrangou, R., C. Fremaux, H. Deveau, M. Richards, P. Boyaval, S. Moineau, D. A. Romero, and P. Horvath. 2007. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709-1712. - PubMed
-
- Beltran, P., J. M. Musser, R. Helmuth, J. J. Farmer III, W. M. Frerichs, I. K. Wachsmuth, K. Ferris, A. C. McWhorter, J. G. Wells, A. Cravioto, and R. K. Selander. 1988. Toward a population genetic analysis of Salmonella: genetic diversity and relationships among strains of serotypes S. choleraesuis, S. derby, S. dublin, S. enteritidis, S. heidelberg, S. infantis, S. newport, and S. typhimurium. Proc. Natl. Acad. Sci. U. S. A. 85:7753-7757. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

