Identification of DysI, the immunity factor of the streptococcal bacteriocin dysgalacticin
- PMID: 20935130
- PMCID: PMC2988609
- DOI: 10.1128/AEM.01707-10
Identification of DysI, the immunity factor of the streptococcal bacteriocin dysgalacticin
Abstract
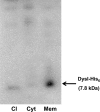
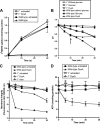
DysI is identified as the protein that confers specific immunity to dysgalacticin, a plasmid-encoded streptococcal bacteriocin. dysI is transcribed as part of the copG-repB-dysI replication-associated operon. DysI appears to function at the membrane level to prevent the inhibitory effects of dysgalacticin on glucose transport, membrane integrity, and intracellular ATP content.
Figures

References
-
- Bendtsen, J. D., H. Nielsen, G. von Heijne, and S. Brunak. 2004. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 340:783-795. - PubMed
-
- Claesson, M. J., Y. Li, S. Leahy, C. Canchaya, J. P. van Pijkeren, A. M. Cerdeño-Tárraga, J. Parkhill, S. Flynn, G. C. O'Sullivan, J. K. Collins, D. Higgins, F. Shanahan, G. F. Fitzgerald, D. van Sinderen, and P. W. O'Toole. 2006. Multireplicon genome architecture of Lactobacillus salivarius. Proc. Natl. Acad. Sci. U. S. A. 103:6718-6723. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

