Comparative structural analysis of human DEAD-box RNA helicases
- PMID: 20941364
- PMCID: PMC2948006
- DOI: 10.1371/journal.pone.0012791
Comparative structural analysis of human DEAD-box RNA helicases
Abstract
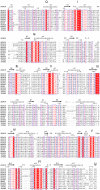
DEAD-box RNA helicases play various, often critical, roles in all processes where RNAs are involved. Members of this family of proteins are linked to human disease, including cancer and viral infections. DEAD-box proteins contain two conserved domains that both contribute to RNA and ATP binding. Despite recent advances the molecular details of how these enzymes convert chemical energy into RNA remodeling is unknown. We present crystal structures of the isolated DEAD-domains of human DDX2A/eIF4A1, DDX2B/eIF4A2, DDX5, DDX10/DBP4, DDX18/myc-regulated DEAD-box protein, DDX20, DDX47, DDX52/ROK1, and DDX53/CAGE, and of the helicase domains of DDX25 and DDX41. Together with prior knowledge this enables a family-wide comparative structural analysis. We propose a general mechanism for opening of the RNA binding site. This analysis also provides insights into the diversity of DExD/H- proteins, with implications for understanding the functions of individual family members.
Conflict of interest statement
Figures





Similar articles
-
Crystal structure of human RNA helicase A (DHX9): structural basis for unselective nucleotide base binding in a DEAD-box variant protein.J Mol Biol. 2010 Jul 23;400(4):768-82. doi: 10.1016/j.jmb.2010.05.046. Epub 2010 May 25. J Mol Biol. 2010. PMID: 20510246
-
Structural basis for RNA-duplex recognition and unwinding by the DEAD-box helicase Mss116p.Nature. 2012 Oct 4;490(7418):121-5. doi: 10.1038/nature11402. Epub 2012 Sep 2. Nature. 2012. PMID: 22940866 Free PMC article.
-
The DEXD/H-box RNA helicase DDX19 is regulated by an {alpha}-helical switch.J Biol Chem. 2009 Apr 17;284(16):10296-300. doi: 10.1074/jbc.C900018200. Epub 2009 Feb 25. J Biol Chem. 2009. PMID: 19244245 Free PMC article.
-
ATP utilization and RNA conformational rearrangement by DEAD-box proteins.Annu Rev Biophys. 2012;41:247-67. doi: 10.1146/annurev-biophys-050511-102243. Epub 2012 Feb 13. Annu Rev Biophys. 2012. PMID: 22404686 Free PMC article. Review.
-
DEAD-box helicases: the Yin and Yang roles in viral infections.Biotechnol Genet Eng Rev. 2018 Apr;34(1):3-32. doi: 10.1080/02648725.2018.1467146. Epub 2018 May 9. Biotechnol Genet Eng Rev. 2018. PMID: 29742983 Review.
Cited by
-
Characterization of the Oncogenic Potential of Eukaryotic Initiation Factor 4A1 in Lung Adenocarcinoma via Cell Cycle Regulation and Immune Microenvironment Reprogramming.Biology (Basel). 2022 Jun 28;11(7):975. doi: 10.3390/biology11070975. Biology (Basel). 2022. PMID: 36101357 Free PMC article.
-
DDX RNA helicases: key players in cellular homeostasis and innate antiviral immunity.J Virol. 2024 Oct 22;98(10):e0004024. doi: 10.1128/jvi.00040-24. Epub 2024 Aug 30. J Virol. 2024. PMID: 39212449 Free PMC article. Review.
-
DEAD/H-Box Helicases in Immunity, Inflammation, Cell Differentiation, and Cell Death and Disease.Cells. 2022 May 11;11(10):1608. doi: 10.3390/cells11101608. Cells. 2022. PMID: 35626643 Free PMC article. Review.
-
Crystal structure, mutational analysis and RNA-dependent ATPase activity of the yeast DEAD-box pre-mRNA splicing factor Prp28.Nucleic Acids Res. 2014 Nov 10;42(20):12885-98. doi: 10.1093/nar/gku930. Epub 2014 Oct 10. Nucleic Acids Res. 2014. PMID: 25303995 Free PMC article.
-
The human Dicer helicase domain is capable of ATP hydrolysis and single-stranded nucleic acid binding.BMC Biol. 2024 Dec 18;22(1):287. doi: 10.1186/s12915-024-02082-x. BMC Biol. 2024. PMID: 39695695 Free PMC article.
References
-
- Linder P, Tanner NK, Banroques J. From RNA helicases to RNPases. Trends Biochem Sci. 2001;26:339–341. - PubMed
-
- Schwer B. A new twist on RNA helicases: DExH/D box proteins as RNPases. Nat Struct Biol. 2001;8:113–116. - PubMed
-
- Cordin O, Banroques J, Tanner NK, Linder P. The DEAD-box protein family of RNA helicases. Gene. 2006;367:17–37. - PubMed
-
- Pyle AM. Translocation and unwinding mechanisms of RNA and DNA helicases. Annu Rev Biophys. 2008;37:317–336. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

