Gene-environment interactions at nucleotide resolution
- PMID: 20941394
- PMCID: PMC2947989
- DOI: 10.1371/journal.pgen.1001144
Gene-environment interactions at nucleotide resolution
Abstract
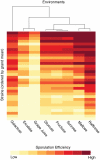
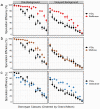
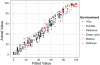
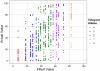
Interactions among genes and the environment are a common source of phenotypic variation. To characterize the interplay between genetics and the environment at single nucleotide resolution, we quantified the genetic and environmental interactions of four quantitative trait nucleotides (QTN) that govern yeast sporulation efficiency. We first constructed a panel of strains that together carry all 32 possible combinations of the 4 QTN genotypes in 2 distinct genetic backgrounds. We then measured the sporulation efficiencies of these 32 strains across 8 controlled environments. This dataset shows that variation in sporulation efficiency is shaped largely by genetic and environmental interactions. We find clear examples of QTN:environment, QTN: background, and environment:background interactions. However, we find no QTN:QTN interactions that occur consistently across the entire dataset. Instead, interactions between QTN only occur under specific combinations of environment and genetic background. Thus, what might appear to be a QTN:QTN interaction in one background and environment becomes a more complex QTN:QTN:environment:background interaction when we consider the entire dataset as a whole. As a result, the phenotypic impact of a set of QTN alleles cannot be predicted from genotype alone. Our results instead demonstrate that the effects of QTN and their interactions are inextricably linked both to genetic background and to environmental variation.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Smith EN, Kruglyak L. Gene-environment interaction in yeast gene expression. PLoS Biol. 2008;6:e83. doi: 10.1371/journal.pbio.0060083. - DOI - PMC - PubMed
-
- Yaffe K, Haan M, Byers A, Tangen C, Kuller L. Estrogen use, APOE, and cognitive decline: evidence of gene-environment interaction. Neurology. 2000;54:1949–1954. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

