Global profiling of proteolysis during rupture of Plasmodium falciparum from the host erythrocyte
- PMID: 20943600
- PMCID: PMC3098579
- DOI: 10.1074/mcp.M110.001636
Global profiling of proteolysis during rupture of Plasmodium falciparum from the host erythrocyte
Abstract
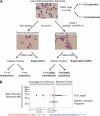
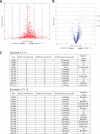
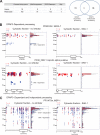
The obligate intracellular parasite pathogen Plasmodium falciparum is the causative agent of malaria, a disease that results in nearly one million deaths per year. A key step in disease pathology in the human host is the parasite-mediated rupture of red blood cells, a process that requires extensive proteolysis of a number of host and parasite proteins. However, only a relatively small number of specific proteolytic processing events have been characterized. Here we describe the application of the Protein Topography and Migration Analysis Platform (PROTOMAP) (Dix, M. M., Simon, G. M., and Cravatt, B. F. (2008) Global mapping of the topography and magnitude of proteolytic events in apoptosis. Cell 134, 679-691; Simon, G. M., Dix, M. M., and Cravatt, B. F. (2009) Comparative assessment of large-scale proteomic studies of apoptotic proteolysis. ACS Chem. Biol. 4, 401-408) technology to globally profile proteolytic events occurring over the last 6-8 h of the intraerythrocytic cycle of P. falciparum. Using this method, we were able to generate peptographs for a large number of proteins at 6 h prior to rupture as well as at the point of rupture and in purified merozoites after exit from the host cell. These peptographs allowed assessment of proteolytic processing as well as changes in both protein localization and overall stage-specific expression of a large number of parasite proteins. Furthermore, by using a highly selective inhibitor of the cysteine protease dipeptidyl aminopeptidase 3 (DPAP3) that has been shown to be a key regulator of host cell rupture, we were able to identify specific substrates whose processing may be of particular importance to the process of host cell rupture. These results provide the first global map of the proteolytic processing events that take place as the human malarial parasite extracts itself from the host red blood cell. These data also provide insight into the biochemical events that take place during host cell rupture and are likely to be valuable for the study of proteases that could potentially be targeted for therapeutic gain.
Figures



References
-
- Arastu-Kapur S., Ponder E. L., Fonoviæ U. P., Yeoh S., Yuan F., Fonoviæ M., Grainger M., Phillips C. I., Powers J. C., Bogyo M. (2008) Identification of proteases that regulate erythrocyte rupture by the malaria parasite Plasmodium falciparum. Nat. Chem. Biol. 4, 203–213 - PubMed
-
- Yeoh S., O'Donnell R. A., Koussis K., Dluzewski A. R., Ansell K. H., Osborne S. A., Hackett F., Withers-Martinez C., Mitchell G. H., Bannister L. H., Bryans J. S., Kettleborough C. A., Blackman M. J. (2007) Subcellular discharge of a serine protease mediates release of invasive malaria parasites from host erythrocytes. Cell 131, 1072–1083 - PubMed
-
- Gevaert K., Goethals M., Martens L., Van Damme J., Staes A., Thomas G. R., Vandekerckhove J. (2003) Exploring proteomes and analyzing protein processing by mass spectrometric identification of sorted N-terminal peptides. Nat. Biotechnol. 21, 566–569 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

