Interaction of poxvirus intracellular mature virion proteins with the TPR domain of kinesin light chain in live infected cells revealed by two-photon-induced fluorescence resonance energy transfer fluorescence lifetime imaging microscopy
- PMID: 20943972
- PMCID: PMC3004322
- DOI: 10.1128/JVI.01395-10
Interaction of poxvirus intracellular mature virion proteins with the TPR domain of kinesin light chain in live infected cells revealed by two-photon-induced fluorescence resonance energy transfer fluorescence lifetime imaging microscopy
Abstract
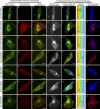
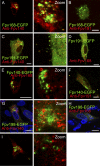
Using two-photon-induced fluorescence lifetime imaging microscopy, we corroborate an interaction (previously demonstrated by yeast two-hybrid domain analysis) of full-length vaccinia virus (VACV; an orthopoxvirus) A36 protein with the cellular microtubule motor protein kinesin. Quenching of enhanced green fluorescent protein (EGFP), fused to the C terminus of VACV A36, by monomeric red fluorescent protein (mDsRed), fused to the tetratricopeptide repeat (TPR) domain of kinesin, was observed in live chicken embryo fibroblasts infected with either modified vaccinia virus Ankara (MVA) or wild-type fowlpox virus (FWPV; an avipoxvirus), and the excited-state fluorescence lifetime of EGFP was reduced from 2.5 ± 0.1 ns to 2.1 ± 0.1 ns due to resonance energy transfer to mDsRed. FWPV does not encode an equivalent of intracellular enveloped virion surface protein A36, yet it is likely that this virus too must interact with kinesin to facilitate intracellular virion transport. To investigate possible interactions between innate FWPV proteins and kinesin, recombinant FWPVs expressing EGFP fused to the N termini of FWPV structural proteins Fpv140, Fpv168, Fpv191, and Fpv198 (equivalent to VACV H3, A4, p4c, and A34, respectively) were generated. EGFP fusions of intracellular mature virion (IMV) surface protein Fpv140 and type II membrane protein Fpv198 were quenched by mDsRed-TPR in recombinant FWPV-infected cells, indicating that these virion proteins are found within 10 nm of mDsRed-TPR. In contrast, and as expected, EGFP fusions of the IMV core protein Fpv168 did not show any quenching. Interestingly, the p4c-like protein Fpv191, which demonstrates late association with preassembled IMV, also did not show any quenching.
Figures


Similar articles
-
Direct interaction of baculovirus capsid proteins VP39 and EXON0 with kinesin-1 in insect cells determined by fluorescence resonance energy transfer-fluorescence lifetime imaging microscopy.J Virol. 2012 Jan;86(2):844-53. doi: 10.1128/JVI.06109-11. Epub 2011 Nov 9. J Virol. 2012. PMID: 22072745 Free PMC article.
-
Vaccinia virus A36R membrane protein provides a direct link between intracellular enveloped virions and the microtubule motor kinesin.J Virol. 2004 Mar;78(5):2486-93. doi: 10.1128/jvi.78.5.2486-2493.2004. J Virol. 2004. PMID: 14963148 Free PMC article.
-
Vaccinia protein F12 has structural similarity to kinesin light chain and contains a motor binding motif required for virion export.PLoS Pathog. 2010 Feb 26;6(2):e1000785. doi: 10.1371/journal.ppat.1000785. PLoS Pathog. 2010. PMID: 20195521 Free PMC article.
-
The exit of vaccinia virus from infected cells.Virus Res. 2004 Dec;106(2):189-97. doi: 10.1016/j.virusres.2004.08.015. Virus Res. 2004. PMID: 15567497 Review.
-
Recent advances using green and red fluorescent protein variants.Appl Microbiol Biotechnol. 2007 Nov;77(1):1-12. doi: 10.1007/s00253-007-1131-5. Epub 2007 Aug 18. Appl Microbiol Biotechnol. 2007. PMID: 17704916 Review.
Cited by
-
Direct interaction of baculovirus capsid proteins VP39 and EXON0 with kinesin-1 in insect cells determined by fluorescence resonance energy transfer-fluorescence lifetime imaging microscopy.J Virol. 2012 Jan;86(2):844-53. doi: 10.1128/JVI.06109-11. Epub 2011 Nov 9. J Virol. 2012. PMID: 22072745 Free PMC article.
-
Identification of giant Mimivirus protein functions using RNA interference.Front Microbiol. 2015 Apr 28;6:345. doi: 10.3389/fmicb.2015.00345. eCollection 2015. Front Microbiol. 2015. PMID: 25972846 Free PMC article.
-
Methodology for the efficient generation of fluorescently tagged vaccinia virus proteins.J Vis Exp. 2014 Jan 17;(83):e51151. doi: 10.3791/51151. J Vis Exp. 2014. PMID: 24473272 Free PMC article.
-
A kinesin-1 binding motif in vaccinia virus that is widespread throughout the human genome.EMBO J. 2011 Nov 16;30(22):4523-38. doi: 10.1038/emboj.2011.326. EMBO J. 2011. PMID: 21915095 Free PMC article.
-
Vaccinia virus proteins A36 and F12/E2 show strong preferences for different kinesin light chain isoforms.Traffic. 2017 Aug;18(8):505-518. doi: 10.1111/tra.12494. Epub 2017 Jun 27. Traffic. 2017. PMID: 28485852 Free PMC article.
References
-
- Boulanger, D., T. Smith, and M. A. Skinner. 2000. Morphogenesis and release of fowlpox virus. J. Gen. Virol. 81:675-687. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

