SIRT3 substrate specificity determined by peptide arrays and machine learning
- PMID: 20945913
- PMCID: PMC3042044
- DOI: 10.1021/cb100218d
SIRT3 substrate specificity determined by peptide arrays and machine learning
Abstract
Accumulating evidence suggests that reversible protein acetylation may be a major regulatory mechanism that rivals phosphorylation. With the recent cataloging of thousands of acetylation sites on hundreds of proteins comes the challenge of identifying the acetyltransferases and deacetylases that regulate acetylation levels. Sirtuins are a conserved family of NAD(+)-dependent protein deacetylases that are implicated in genome maintenance, metabolism, cell survival, and lifespan. SIRT3 is the dominant protein deacetylase in mitochondria, and emerging evidence suggests that SIRT3 may control major pathways by deacetylation of central metabolic enzymes. Here, to identify potential SIRT3 substrates, we have developed an unbiased screening strategy that involves a novel acetyl-lysine analogue (thiotrifluoroacetyl-lysine), SPOT-peptide libraries, machine learning, and kinetic validation. SPOT peptide libraries based on known and potential mitochondrial acetyl-lysine sites were screened for SIRT3 binding and then analyzed using machine learning to establish binding trends. These trends were then applied to the mitochondrial proteome as a whole to predict binding affinity of all lysine sites within human mitochondria. Machine learning prediction of SIRT3 binding correlated with steady-state kinetic k(cat)/K(m) values for 24 acetyl-lysine peptides that possessed a broad range of predicted binding. Thus, SPOT peptide-binding screens and machine learning prediction provides an accurate and efficient method to evaluate sirtuin substrate specificity from a relatively small learning set. These analyses suggest potential SIRT3 substrates involved in several metabolic pathways such as the urea cycle, ATP synthesis, and fatty acid oxidation.
Figures




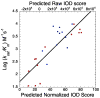

References
-
- Kim SC, Sprung R, Chen Y, Xu Y, Ball H, Pei J, Cheng T, Kho Y, Xiao H, Xiao L, Grishin NV, White M, Yang XJ, Zhao Y. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol Cell. 2006;23:607–618. - PubMed
-
- Choudhary C, Kumar C, Gnad F, Nielsen ML, Rehman M, Walther TC, Olsen JV, Mann M. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science. 2009;325:834–840. - PubMed
-
- Longo VD, Kennedy BK. Sirtuins in aging and age-related disease. Cell. 2006;126:257–268. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

