Association of mouse Dlg4 (PSD-95) gene deletion and human DLG4 gene variation with phenotypes relevant to autism spectrum disorders and Williams' syndrome
- PMID: 20952458
- PMCID: PMC3146008
- DOI: 10.1176/appi.ajp.2010.10040484
Association of mouse Dlg4 (PSD-95) gene deletion and human DLG4 gene variation with phenotypes relevant to autism spectrum disorders and Williams' syndrome
Abstract
Objective: Research is increasingly linking autism spectrum disorders and other neurodevelopmental disorders to synaptic abnormalities ("synaptopathies"). PSD-95 (postsynaptic density-95, DLG4) orchestrates protein-protein interactions at excitatory synapses and is a major functional bridge interconnecting a neurexinneuroligin-SHANK pathway implicated in autism spectrum disorders.
Method: The authors characterized behavioral, dendritic, and molecular phenotypic abnormalities relevant to autism spectrum disorders in mice with PSD-95 deletion (Dlg4⁻(/)⁻). The data from mice led to the identification of single-nucleotide polymorphisms (SNPs) in human DLG4 and the examination of associations between these variants and neural signatures of Williams' syndrome in a normal population, using functional and structural neuroimaging.
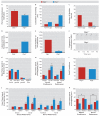
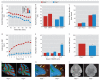
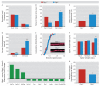
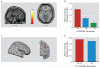
Results: Dlg4⁻(/)⁻ showed increased repetitive behaviors, abnormal communication and social behaviors, impaired motor coordination, and increased stress reactivity and anxiety-related responses. Dlg4⁻(/)⁻ had subtle dysmorphology of amygdala dendritic spines and altered forebrain expression of various synaptic genes, including Cyln2, which regulates cytoskeletal dynamics and is a candidate gene for Williams' syndrome. A signifi-cant association was observed between variations in two human DLG4 SNPs and reduced intraparietal sulcus volume and abnormal cortico-amygdala coupling, both of which characterize Williams' syndrome.
Conclusions: These findings demonstrate that DLG4 gene disruption in mice produces a complex range of behavioral and molecular abnormalities relevant to autism spectrum disorders and Williams' syndrome. The study provides an initial link between human DLG4 gene variation and key neural endophenotypes of Williams' syndrome and perhaps corticoamygdala regulation of emotional and social processes more generally.
Figures
Similar articles
-
Glutamatergic postsynaptic density protein dysfunctions in synaptic plasticity and dendritic spines morphology: relevance to schizophrenia and other behavioral disorders pathophysiology, and implications for novel therapeutic approaches.Mol Neurobiol. 2014 Feb;49(1):484-511. doi: 10.1007/s12035-013-8534-3. Epub 2013 Sep 3. Mol Neurobiol. 2014. PMID: 23999870 Review.
-
Population-specific haplotype association of the postsynaptic density gene DLG4 with schizophrenia, in family-based association studies.PLoS One. 2013 Jul 25;8(7):e70302. doi: 10.1371/journal.pone.0070302. Print 2013. PLoS One. 2013. PMID: 23936182 Free PMC article.
-
Cerebellar Shank2 Regulates Excitatory Synapse Density, Motor Coordination, and Specific Repetitive and Anxiety-Like Behaviors.J Neurosci. 2016 Nov 30;36(48):12129-12143. doi: 10.1523/JNEUROSCI.1849-16.2016. J Neurosci. 2016. PMID: 27903723 Free PMC article.
-
Association of GTF2i in the Williams-Beuren syndrome critical region with autism spectrum disorders.J Autism Dev Disord. 2012 Jul;42(7):1459-69. doi: 10.1007/s10803-011-1389-4. J Autism Dev Disord. 2012. PMID: 22048961
-
Conceptualizing neurodevelopmental disorders through a mechanistic understanding of fragile X syndrome and Williams syndrome.Curr Opin Neurol. 2012 Apr;25(2):112-24. doi: 10.1097/WCO.0b013e328351823c. Curr Opin Neurol. 2012. PMID: 22395002 Free PMC article. Review.
Cited by
-
Synapse and Receptor Alterations in Two Different S100B-Induced Glaucoma-Like Models.Int J Mol Sci. 2020 Sep 23;21(19):6998. doi: 10.3390/ijms21196998. Int J Mol Sci. 2020. PMID: 32977518 Free PMC article.
-
Expression of the G72/G30 gene in transgenic mice induces behavioral changes.Mol Psychiatry. 2014 Feb;19(2):175-83. doi: 10.1038/mp.2012.185. Epub 2013 Jan 22. Mol Psychiatry. 2014. PMID: 23337943 Free PMC article.
-
Altered microstructure within social-cognitive brain networks during childhood in Williams syndrome.Cereb Cortex. 2014 Oct;24(10):2796-806. doi: 10.1093/cercor/bht135. Epub 2013 May 24. Cereb Cortex. 2014. PMID: 23709644 Free PMC article.
-
Synaptic proteins and receptors defects in autism spectrum disorders.Front Cell Neurosci. 2014 Sep 11;8:276. doi: 10.3389/fncel.2014.00276. eCollection 2014. Front Cell Neurosci. 2014. PMID: 25309321 Free PMC article. Review.
-
Tetramethylpyrazine nitrone activates the BDNF/Akt/CREB pathway to promote post-ischaemic neuroregeneration and recovery of neurological functions in rats.Br J Pharmacol. 2018 Feb;175(3):517-531. doi: 10.1111/bph.14102. Epub 2017 Dec 22. Br J Pharmacol. 2018. PMID: 29161771 Free PMC article.
References
-
- Marshall CR, Noor A, Vincent JB, Lionel AC, Feuk L, Skaug J, Shago M, Moessner R, Pinto D, Ren Y, Thiruvahindrapduram B, Fiebig A, Schreiber S, Friedman J, Ketelaars CE, Vos YJ, Ficicioglu C, Kirkpatrick S, Nicolson R, Sloman L, Summers A, Gibbons CA, Teebi A, Chitayat D, Weksberg R, Thompson A, Vardy C, Crosbie V, Luscombe S, Baatjes R, Zwaigenbaum L, Roberts W, Fernandez B, Szatmari P, Scherer SW. Structural variation of chromosomes in autism spectrum disorder. Am J Hum Genet. 2008;82:477–488. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

