Binding-induced folding of prokaryotic ubiquitin-like protein on the Mycobacterium proteasomal ATPase targets substrates for degradation
- PMID: 20953180
- PMCID: PMC2988878
- DOI: 10.1038/nsmb.1918
Binding-induced folding of prokaryotic ubiquitin-like protein on the Mycobacterium proteasomal ATPase targets substrates for degradation
Abstract
Mycobacterium tuberculosis uses a proteasome system that is analogous to the eukaryotic ubiquitin-proteasome pathway and is required for pathogenesis. However, the bacterial analog of ubiquitin, prokaryotic ubiquitin-like protein (Pup), is an intrinsically disordered protein that bears little sequence or structural resemblance to the highly structured ubiquitin. Thus, it was unknown how pupylated proteins were recruited to the proteasome. Here, we show that the Mycobacterium proteasomal ATPase (Mpa) has three pairs of tentacle-like coiled coils that recognize Pup. Mpa bound unstructured Pup through hydrophobic interactions and a network of hydrogen bonds, leading to the formation of an α-helix in Pup. Our work describes a binding-induced folding recognition mechanism in the Pup-proteasome system that differs mechanistically from substrate recognition in the ubiquitin-proteasome system. This key difference between the prokaryotic and eukaryotic systems could be exploited for the development of a small molecule-based treatment for tuberculosis.
Figures


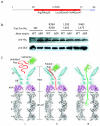
Similar articles
-
The mycobacterial Mpa-proteasome unfolds and degrades pupylated substrates by engaging Pup's N-terminus.EMBO J. 2010 Apr 7;29(7):1262-71. doi: 10.1038/emboj.2010.23. Epub 2010 Mar 4. EMBO J. 2010. PMID: 20203624 Free PMC article.
-
Prokaryotic ubiquitin-like protein remains intrinsically disordered when covalently attached to proteasomal target proteins.BMC Struct Biol. 2017 Feb 1;17(1):1. doi: 10.1186/s12900-017-0072-1. BMC Struct Biol. 2017. PMID: 28143508 Free PMC article.
-
Prokaryotic ubiquitin-like protein pup is intrinsically disordered.J Mol Biol. 2009 Sep 11;392(1):208-17. doi: 10.1016/j.jmb.2009.07.018. Epub 2009 Jul 14. J Mol Biol. 2009. PMID: 19607839 Free PMC article.
-
Pupylation as a signal for proteasomal degradation in bacteria.Biochim Biophys Acta. 2014 Jan;1843(1):103-13. doi: 10.1016/j.bbamcr.2013.03.022. Epub 2013 Apr 2. Biochim Biophys Acta. 2014. PMID: 23557784 Review.
-
Targets of ubiquitin like system in mycobacteria and related actinobacterial species.Microbiol Res. 2017 Nov;204:9-29. doi: 10.1016/j.micres.2017.07.002. Epub 2017 Jul 18. Microbiol Res. 2017. PMID: 28870295 Review.
Cited by
-
The pup-proteasome system of Mycobacterium tuberculosis.Subcell Biochem. 2013;66:267-95. doi: 10.1007/978-94-007-5940-4_10. Subcell Biochem. 2013. PMID: 23479444 Free PMC article. Review.
-
The β-Grasp Domain of Proteasomal ATPase Mpa Makes Critical Contacts with the Mycobacterium tuberculosis 20S Core Particle to Facilitate Degradation.mSphere. 2022 Oct 26;7(5):e0027422. doi: 10.1128/msphere.00274-22. Epub 2022 Aug 22. mSphere. 2022. PMID: 35993699 Free PMC article.
-
Characterization of the bipartite degron that regulates ubiquitin-independent degradation of thymidylate synthase.Biosci Rep. 2013 Jan 18;33(1):165-73. doi: 10.1042/BSR20120112. Biosci Rep. 2013. PMID: 23181752 Free PMC article.
-
Fluorescent probes reveal a minimal ligase recognition motif in the prokaryotic ubiquitin-like protein from Mycobacterium tuberculosis.J Am Chem Soc. 2013 Feb 27;135(8):2887-90. doi: 10.1021/ja311376h. Epub 2013 Feb 14. J Am Chem Soc. 2013. PMID: 23402667 Free PMC article.
-
Allosteric transitions direct protein tagging by PafA, the prokaryotic ubiquitin-like protein (Pup) ligase.J Biol Chem. 2013 Apr 19;288(16):11287-93. doi: 10.1074/jbc.M112.435842. Epub 2013 Mar 7. J Biol Chem. 2013. PMID: 23471967 Free PMC article.
References
-
- Lupas A, et al. Eubacterial proteasomes. Mol Biol Rep. 1997;24:125–31. - PubMed
-
- Lin G, et al. Mycobacterium tuberculosis prcBA genes encode a gated proteasome with broad oligopeptide specificity. Mol Microbiol. 2006;59:1405–16. - PubMed
-
- Hu G, et al. Structure of the Mycobacterium tuberculosis proteasome and mechanism of inhibition by a peptidyl boronate. Mol Microbiol. 2006;59:1417–28. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

