Dimerization of protegrin-1 in different environments
- PMID: 20957087
- PMCID: PMC2956088
- DOI: 10.3390/ijms11093177
Dimerization of protegrin-1 in different environments
Abstract
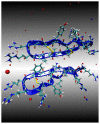
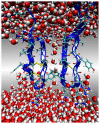
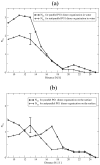
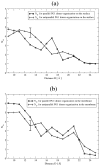
The dimerization of the cationic β-hairpin antimicrobial peptide protegrin-1 (PG1) is investigated in three different environments: water, the surface of a lipid bilayer membrane, and the core of the membrane. PG1 is known to kill bacteria by forming oligomeric membrane pores, which permeabilize the cells. PG1 dimers are found in two distinct, parallel and antiparallel, conformations, known as important intermediate structural units of the active pore oligomers. What is not clear is the sequence of events from PG1 monomers in solution to pores inside membranes. The step we focus on in this work is the dimerization of PG1. In particular, we are interested in determining where PG1 dimerization is most favorable. We use extensive molecular dynamics simulations to determine the potential of mean force as a function of distance between two PG1 monomers in the aqueous subphase, the surface of model lipid bilayers and the interior of these bilayers. We investigate the two known distinct modes of dimerization that result in either a parallel or an antiparallel β-sheet orientation. The model bilayer membranes are composed of anionic palmitoyl-oleoyl-phosphatidylglycerol (POPG) and palmitoyl-oleoyl-phosphatidylethanolamine (POPE) in a 1:3 ratio (POPG:POPE). We find the parallel PG1 dimer association to be more favorable than the antiparallel one in water and inside the membrane. However, we observe that the antiparallel PG1 β-sheet dimer conformation is somewhat more stable than the parallel dimer association at the surface of the membrane. We explore the role of hydrogen bonds and ionic bridges in peptide dimerization in the three environments. Detailed knowledge of how networks of ionic bridges and hydrogen bonds contribute to peptide stability is essential for the purpose of understanding the mechanism of action for membrane-active peptides as well as for designing peptides which can modulate membrane properties. The findings are suggestive of the dominant pathways leading from individual PG1 molecules in solution to functional pores in bacterial membranes.
Keywords: dimerization; potential of mean force; protegrin.
Figures


Similar articles
-
Computational studies of protegrin antimicrobial peptides: a review.Peptides. 2011 Jan;32(1):188-201. doi: 10.1016/j.peptides.2010.10.006. Epub 2010 Oct 12. Peptides. 2011. PMID: 20946928 Free PMC article. Review.
-
Free energy profile of the interaction between a monomer or a dimer of protegrin-1 in a specific binding orientation and a model lipid bilayer.J Phys Chem B. 2010 Mar 4;114(8):2790-7. doi: 10.1021/jp909640g. J Phys Chem B. 2010. PMID: 20136112 Free PMC article.
-
Conformational study of the protegrin-1 (PG-1) dimer interaction with lipid bilayers and its effect.BMC Struct Biol. 2007 Apr 2;7:21. doi: 10.1186/1472-6807-7-21. BMC Struct Biol. 2007. PMID: 17407565 Free PMC article.
-
Membrane interactions and pore formation by the antimicrobial peptide protegrin.Biophys J. 2013 Feb 5;104(3):633-42. doi: 10.1016/j.bpj.2012.12.038. Biophys J. 2013. PMID: 23442914 Free PMC article.
-
Biological activity and structural aspects of PGLa interaction with membrane mimetic systems.Biochim Biophys Acta. 2009 Aug;1788(8):1656-66. doi: 10.1016/j.bbamem.2009.05.012. Epub 2009 May 29. Biochim Biophys Acta. 2009. PMID: 19481533 Review.
Cited by
-
Computational prediction of the optimal oligomeric state for membrane-inserted β-barrels of protegrin-1 and related mutants.J Pept Sci. 2017 Apr;23(4):334-345. doi: 10.1002/psc.2992. J Pept Sci. 2017. PMID: 28382709 Free PMC article.
-
Computational studies of protegrin antimicrobial peptides: a review.Peptides. 2011 Jan;32(1):188-201. doi: 10.1016/j.peptides.2010.10.006. Epub 2010 Oct 12. Peptides. 2011. PMID: 20946928 Free PMC article. Review.
-
Multiscale models of the antimicrobial peptide protegrin-1 on gram-negative bacteria membranes.Int J Mol Sci. 2012;13(9):11000-11011. doi: 10.3390/ijms130911000. Epub 2012 Sep 5. Int J Mol Sci. 2012. PMID: 23109834 Free PMC article. Review.
-
Oligomerization of the antimicrobial peptide Protegrin-5 in a membrane-mimicking environment. Structural studies by high-resolution NMR spectroscopy.Eur Biophys J. 2017 Apr;46(3):293-300. doi: 10.1007/s00249-016-1167-5. Epub 2016 Sep 2. Eur Biophys J. 2017. PMID: 27589857
-
Computational studies of peptide-induced membrane pore formation.Philos Trans R Soc Lond B Biol Sci. 2017 Aug 5;372(1726):20160219. doi: 10.1098/rstb.2016.0219. Philos Trans R Soc Lond B Biol Sci. 2017. PMID: 28630158 Free PMC article. Review.
References
-
- Fahrner RL, Dieckmann T, Harwig SS, Lehrer RI, Eisenberg D, Feigon J. Solution structure of protegrin-1, a broad-spectrum antimicrobial peptide from porcine leukocytes. Chem. Biol. 1996;3:543–550. - PubMed
-
- Mani R, Tang M, Wu X, Buffy IJ, Waring AJ, Sherman MA, Hong M. Membrane-bound dimer structure of a beta-hairpin antimicrobial peptide from rotational-echo double-resonance solid state NMR. Biochemistry. 2006;45:8341–8349. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

