Ray: simultaneous assembly of reads from a mix of high-throughput sequencing technologies
- PMID: 20958248
- PMCID: PMC3119603
- DOI: 10.1089/cmb.2009.0238
Ray: simultaneous assembly of reads from a mix of high-throughput sequencing technologies
Abstract
An accurate genome sequence of a desired species is now a pre-requisite for genome research. An important step in obtaining a high-quality genome sequence is to correctly assemble short reads into longer sequences accurately representing contiguous genomic regions. Current sequencing technologies continue to offer increases in throughput, and corresponding reductions in cost and time. Unfortunately, the benefit of obtaining a large number of reads is complicated by sequencing errors, with different biases being observed with each platform. Although software are available to assemble reads for each individual system, no procedure has been proposed for high-quality simultaneous assembly based on reads from a mix of different technologies. In this paper, we describe a parallel short-read assembler, called Ray, which has been developed to assemble reads obtained from a combination of sequencing platforms. We compared its performance to other assemblers on simulated and real datasets. We used a combination of Roche/454 and Illumina reads to assemble three different genomes. We showed that mixing sequencing technologies systematically reduces the number of contigs and the number of errors. Because of its open nature, this new tool will hopefully serve as a basis to develop an assembler that can be of universal utilization (availability: http://deNovoAssembler.sf.Net/). For online Supplementary Material , see www.liebertonline.com.
Figures

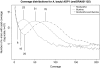

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

