Differential subcellular recruitment of monoacylglycerol lipase generates spatial specificity of 2-arachidonoyl glycerol signaling during axonal pathfinding
- PMID: 20962221
- PMCID: PMC2987617
- DOI: 10.1523/JNEUROSCI.2126-10.2010
Differential subcellular recruitment of monoacylglycerol lipase generates spatial specificity of 2-arachidonoyl glycerol signaling during axonal pathfinding
Abstract
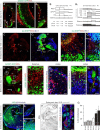
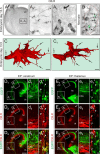
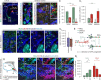
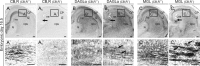
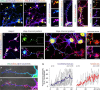
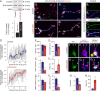
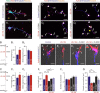
Endocannabinoids, particularly 2-arachidonoyl glycerol (2-AG), impact the directional turning and motility of a developing axon by activating CB(1) cannabinoid receptors (CB(1)Rs) in its growth cone. Recent findings posit that sn-1-diacylglycerol lipases (DAGLα/β) synthesize 2-AG in the motile axon segment of developing pyramidal cells. Coincident axonal targeting of CB(1)Rs and DAGLs prompts the hypothesis that autocrine 2-AG signaling facilitates axonal outgrowth. However, DAGLs alone are insufficient to account for the spatial specificity and dynamics of 2-AG signaling. Therefore, we hypothesized that local 2-AG degradation by monoacylglycerol lipase (MGL) must play a role. We determined how subcellular recruitment of MGL is temporally and spatially restricted to establish the signaling competence of 2-AG during axonal growth. MGL is expressed in central and peripheral axons of the fetal nervous system by embryonic day 12.5. MGL coexists with DAGLα and CB(1)Rs in corticofugal axons of pyramidal cells. Here, MGL and DAGLα undergo differential axonal targeting with MGL being excluded from the motile neurite tip. Thus, spatially confined MGL activity generates a 2-AG-sensing microdomain and configures 2-AG signaling to promote axonal growth. Once synaptogenesis commences, MGL disperses in stationary growth cones. The axonal polarity of MGL is maintained by differential proteasomal degradation because inhibiting the ubiquitin proteasome system also induces axonal MGL redistribution. Because MGL inactivation drives a CB(1)R-dependent axonal growth response, we conclude that 2-AG may act as a focal protrusive signal for developing neurons and whose regulated metabolism is critical for attaining correct axonal complexity.
Figures


Similar articles
-
Nerve growth factor scales endocannabinoid signaling by regulating monoacylglycerol lipase turnover in developing cholinergic neurons.Proc Natl Acad Sci U S A. 2013 Jan 29;110(5):1935-40. doi: 10.1073/pnas.1212563110. Epub 2013 Jan 14. Proc Natl Acad Sci U S A. 2013. PMID: 23319656 Free PMC article.
-
Presynaptic monoacylglycerol lipase activity determines basal endocannabinoid tone and terminates retrograde endocannabinoid signaling in the hippocampus.J Neurosci. 2007 Jan 31;27(5):1211-9. doi: 10.1523/JNEUROSCI.4159-06.2007. J Neurosci. 2007. PMID: 17267577 Free PMC article.
-
Localization of diacylglycerol lipase-alpha around postsynaptic spine suggests close proximity between production site of an endocannabinoid, 2-arachidonoyl-glycerol, and presynaptic cannabinoid CB1 receptor.J Neurosci. 2006 May 3;26(18):4740-51. doi: 10.1523/JNEUROSCI.0054-06.2006. J Neurosci. 2006. PMID: 16672646 Free PMC article.
-
Endocannabinoid overload.Mol Pharmacol. 2010 Dec;78(6):993-5. doi: 10.1124/mol.110.069427. Epub 2010 Oct 15. Mol Pharmacol. 2010. PMID: 20952498 Free PMC article. Review.
-
Biosynthesis and degradation of the endocannabinoid 2-arachidonoylglycerol.Biofactors. 2011 Jan-Feb;37(1):1-7. doi: 10.1002/biof.131. Epub 2010 Nov 29. Biofactors. 2011. PMID: 21328621 Review.
Cited by
-
Endocannabinoids regulate the migration of subventricular zone-derived neuroblasts in the postnatal brain.J Neurosci. 2011 Mar 16;31(11):4000-11. doi: 10.1523/JNEUROSCI.5483-10.2011. J Neurosci. 2011. PMID: 21411643 Free PMC article.
-
A secretagogin locus of the mammalian hypothalamus controls stress hormone release.EMBO J. 2015 Jan 2;34(1):36-54. doi: 10.15252/embj.201488977. Epub 2014 Nov 27. EMBO J. 2015. PMID: 25430741 Free PMC article.
-
Molecular reorganization of endocannabinoid signalling in Alzheimer's disease.Brain. 2011 Apr;134(Pt 4):1041-60. doi: 10.1093/brain/awr046. Brain. 2011. PMID: 21459826 Free PMC article.
-
Marijuana, Spice 'herbal high', and early neural development: implications for rescheduling and legalization.Drug Test Anal. 2013 Jan;5(1):27-45. doi: 10.1002/dta.1390. Epub 2012 Aug 13. Drug Test Anal. 2013. PMID: 22887867 Free PMC article. Review.
-
Parsing the players: 2-arachidonoylglycerol synthesis and degradation in the CNS.Br J Pharmacol. 2014 Mar;171(6):1379-91. doi: 10.1111/bph.12411. Br J Pharmacol. 2014. PMID: 24102242 Free PMC article. Review.
References
-
- Aguado T, Monory K, Palazuelos J, Stella N, Cravatt B, Lutz B, Marsicano G, Kokaia Z, Guzmán M, Galve-Roperh I. The endocannabinoid system drives neural progenitor proliferation. FASEB J. 2005;19:1704–1706. - PubMed
-
- Berghuis P, Rajnicek AM, Morozov YM, Ross RA, Mulder J, Urbán GM, Monory K, Marsicano G, Matteoli M, Canty A, Irving AJ, Katona I, Yanagawa Y, Rakic P, Lutz B, Mackie K, Harkany T. Hardwiring the brain: endocannabinoids shape neuronal connectivity. Science. 2007;316:1212–1216. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
