Autophagy in protists
- PMID: 20962583
- PMCID: PMC3039767
- DOI: 10.4161/auto.7.2.13310
Autophagy in protists
Abstract
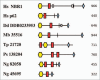
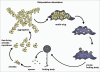
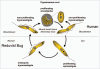
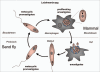
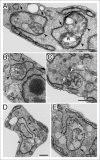
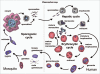
Autophagy is the degradative process by which eukaryotic cells digest their own components using acid hydrolases within the lysosome. Originally thought to function almost exclusively in providing starving cells with nutrients taken from their own cellular constituents, autophagy is in fact involved in numerous cellular events including differentiation, turnover of macromolecules and organelles, and defense against parasitic invaders. During the last 10-20 years, molecular components of the autophagic machinery have been discovered, revealing a complex interactome of proteins and lipids, which, in a concerted way, induce membrane formation to engulf cellular material and target it for lysosomal degradation. Here, our emphasis is autophagy in protists. We discuss experimental and genomic data indicating that the canonical autophagy machinery characterized in animals and fungi appeared prior to the radiation of major eukaryotic lineages. Moreover, we describe how comparative bioinformatics revealed that this canonical machinery has been subject to moderation, outright loss or elaboration on multiple occasions in protist lineages, most probably as a consequence of diverse lifestyle adaptations. We also review experimental studies illustrating how several pathogenic protists either utilize autophagy mechanisms or manipulate host-cell autophagy in order to establish or maintain infection within a host. The essentiality of autophagy for the pathogenicity of many parasites, and the unique features of some of the autophagy-related proteins involved, suggest possible new targets for drug discovery. Further studies of the molecular details of autophagy in protists will undoubtedly enhance our understanding of the diversity and complexity of this cellular phenomenon and the opportunities it offers as a drug target.
Figures





References
-
- de Duve C. The lysosome turns fifty. Nat Cell Biol. 2005;7:847–849. - PubMed
-
- de Reuck AVS, Cameron MP. Ciba Foundation Symposium on Lysosomes. London: J.A. Chrurchill Ltd; 1963.
-
- Klionsky DJ. Autophagy revisited: a conversation with Christian de Duve. Autophagy. 2008;4:740–743. - PubMed
-
- Klionsky DJ. Autophagy: from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol. 2007;8:931–937. - PubMed
-
- Klionsky DJ, Cregg JM, Dunn WA, Jr, Emr SD, Sakai Y, Sandoval IV, et al. A unified nomenclature for yeast autophagy-related genes. Dev Cell. 2003;5:539–545. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
