BriX: a database of protein building blocks for structural analysis, modeling and design
- PMID: 20972210
- PMCID: PMC3013806
- DOI: 10.1093/nar/gkq972
BriX: a database of protein building blocks for structural analysis, modeling and design
Abstract
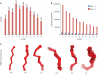
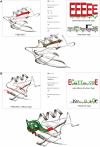
High-resolution structures of proteins remain the most valuable source for understanding their function in the cell and provide leads for drug design. Since the availability of sufficient protein structures to tackle complex problems such as modeling backbone moves or docking remains a problem, alternative approaches using small, recurrent protein fragments have been employed. Here we present two databases that provide a vast resource for implementing such fragment-based strategies. The BriX database contains fragments from over 7000 non-homologous proteins from the Astral collection, segmented in lengths from 4 to 14 residues and clustered according to structural similarity, summing up to a content of 2 million fragments per length. To overcome the lack of loops classified in BriX, we constructed the Loop BriX database of non-regular structure elements, clustered according to end-to-end distance between the regular residues flanking the loop. Both databases are available online (http://brix.crg.es) and can be accessed through a user-friendly web-interface. For high-throughput queries a web-based API is provided, as well as full database downloads. In addition, two exciting applications are provided as online services: (i) user-submitted structures can be covered on the fly with BriX classes, representing putative structural variation throughout the protein and (ii) gaps or low-confidence regions in these structures can be bridged with matching fragments.
Figures
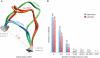

Similar articles
-
Reconstruction of protein backbones from the BriX collection of canonical protein fragments.PLoS Comput Biol. 2008 May 23;4(5):e1000083. doi: 10.1371/journal.pcbi.1000083. PLoS Comput Biol. 2008. PMID: 18483555 Free PMC article.
-
Dynameomics: data-driven methods and models for utilizing large-scale protein structure repositories for improving fragment-based loop prediction.Protein Sci. 2014 Nov;23(11):1584-95. doi: 10.1002/pro.2537. Epub 2014 Sep 3. Protein Sci. 2014. PMID: 25142412 Free PMC article.
-
SSEP: Secondary structural elements of proteins.Nucleic Acids Res. 2003 Jul 1;31(13):3404-5. doi: 10.1093/nar/gkg507. Nucleic Acids Res. 2003. PMID: 12824336 Free PMC article.
-
Protein design with fragment databases.Curr Opin Struct Biol. 2011 Aug;21(4):452-9. doi: 10.1016/j.sbi.2011.05.002. Epub 2011 Jun 16. Curr Opin Struct Biol. 2011. PMID: 21684149 Review.
-
High-throughput modeling and analysis of protein structural dynamics.Brief Bioinform. 2007 Nov;8(6):432-45. doi: 10.1093/bib/bbm014. Epub 2007 May 7. Brief Bioinform. 2007. PMID: 17485424 Review.
Cited by
-
Critical Features of Fragment Libraries for Protein Structure Prediction.PLoS One. 2017 Jan 13;12(1):e0170131. doi: 10.1371/journal.pone.0170131. eCollection 2017. PLoS One. 2017. PMID: 28085928 Free PMC article.
-
Sequence statistics of tertiary structural motifs reflect protein stability.PLoS One. 2017 May 26;12(5):e0178272. doi: 10.1371/journal.pone.0178272. eCollection 2017. PLoS One. 2017. PMID: 28552940 Free PMC article.
-
Secondary structure, a missing component of sequence-based minimotif definitions.PLoS One. 2012;7(12):e49957. doi: 10.1371/journal.pone.0049957. Epub 2012 Dec 7. PLoS One. 2012. PMID: 23236358 Free PMC article.
-
Adaption of human antibody λ and κ light chain architectures to CDR repertoires.Protein Eng Des Sel. 2019 Dec 13;32(3):109-127. doi: 10.1093/protein/gzz012. Protein Eng Des Sel. 2019. PMID: 31535139 Free PMC article.
-
Cryo-EM structure of a lysozyme-derived amyloid fibril from hereditary amyloidosis.Nat Commun. 2024 Nov 7;15(1):9648. doi: 10.1038/s41467-024-54091-7. Nat Commun. 2024. PMID: 39511224 Free PMC article.
References
-
- Kopp J, Bordoli L, Battey JND, Kiefer F, Schwede T. Assessment of CASP7 predictions for template-based modeling targets. Proteins. 2007;69(Suppl. 8):38–56. - PubMed
-
- Murzin AG, Brenner SE, Hubbard T, Chothia C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 1995;247:536–540. - PubMed
-
- Orengo CA, Michie AD, Jones S, Jones DT, Swindells MB, Thornton JM. CATH – a hierarchic classification of protein domain structures. Structure. 1997;5:1093–1108. - PubMed
-
- Fitzkee NC, Fleming PJ, Gong H, Panasik N, Street TO, Rose GD. Are proteins made from a limited parts list? Trends Biochem. Sci. 2005;30:73–80. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

