Early cytokinin response proteins and phosphoproteins of Arabidopsis thaliana identified by proteome and phosphoproteome profiling
- PMID: 20974740
- PMCID: PMC3022391
- DOI: 10.1093/jxb/erq322
Early cytokinin response proteins and phosphoproteins of Arabidopsis thaliana identified by proteome and phosphoproteome profiling
Abstract
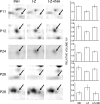
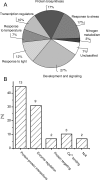
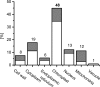
Cytokinins are plant hormones involved in regulation of diverse developmental and physiological processes in plants whose molecular mechanisms of action are being intensely researched. However, most rapid responses to cytokinin signals at the proteomic and phosphoproteomic levels are unknown. Early cytokinin responses were investigated through proteome-wide expression profiling based on image and mass spectrometric analysis of two-dimensionally separated proteins and phosphoproteins. The effects of 15 min treatments of 7-day-old Arabidopsis thaliana seedlings with four main cytokinins representing hydroxyisopentenyl, isopentenyl, aromatic, and urea-derived type cytokinins were compared to help elucidate their common and specific function(s) in regulating plant development. In proteome and phosphoproteome maps, significant differences were reproducibly observed for 53 and 31 protein spots, respectively. In these spots, 96 proteins were identified by matrix-assisted laser desorption/ionization time-of-flight/time-of-flight mass spectrometry (MALDI-TOF/TOF MS), providing a snapshot of early links in cytokinin-regulated signalling circuits and cellular processes, including light signalling and photosynthesis, nitrogen metabolism, the CLAVATA pathway, and protein and gene expression regulation, in accordance with previously described cytokinin functions. Furthermore, they indicate novel links between temperature and cytokinin signalling, and an involvement of calcium ions in cytokinin signalling. Most of the differentially regulated proteins and phosphoproteins are located in chloroplasts, suggesting an as yet uncharacterized direct signalling chain responsible for cytokinin action in chloroplasts. Finally, first insights into the degree of specificity of cytokinin receptors on phosphoproteomic effects were obtained from analyses of cytokinin action in a set of cytokinin receptor double mutants.
Figures


Similar articles
-
Proteome and metabolome profiling of cytokinin action in Arabidopsis identifying both distinct and similar responses to cytokinin down- and up-regulation.J Exp Bot. 2013 Nov;64(14):4193-206. doi: 10.1093/jxb/ert227. Epub 2013 Sep 24. J Exp Bot. 2013. PMID: 24064926 Free PMC article.
-
Cytokinin modulates proteomic, transcriptomic and growth responses to temperature shocks in Arabidopsis.Plant Cell Environ. 2014 Jul;37(7):1641-55. doi: 10.1111/pce.12270. Epub 2014 Feb 13. Plant Cell Environ. 2014. PMID: 24393122
-
Rapid alteration of the phosphoproteome in the moss Physcomitrella patens after cytokinin treatment.J Proteome Res. 2006 Sep;5(9):2283-93. doi: 10.1021/pr060152e. J Proteome Res. 2006. PMID: 16944940
-
Generic signal-specific responses: cytokinin and context-dependent cellular responses.J Exp Bot. 2011 Jun;62(10):3273-88. doi: 10.1093/jxb/erq420. Epub 2011 Jan 6. J Exp Bot. 2011. PMID: 21212299 Review.
-
Cytokinin signal transduction in plant cells.J Plant Res. 2003 Jun;116(3):221-31. doi: 10.1007/s10265-003-0094-6. Epub 2003 Apr 17. J Plant Res. 2003. PMID: 12836044 Review.
Cited by
-
Histidine kinase inhibitors impair shoot regeneration in Arabidopsis thaliana via cytokinin signaling and SAM patterning determinants.Front Plant Sci. 2022 Sep 8;13:894208. doi: 10.3389/fpls.2022.894208. eCollection 2022. Front Plant Sci. 2022. PMID: 36684719 Free PMC article.
-
Proteome and metabolome profiling of cytokinin action in Arabidopsis identifying both distinct and similar responses to cytokinin down- and up-regulation.J Exp Bot. 2013 Nov;64(14):4193-206. doi: 10.1093/jxb/ert227. Epub 2013 Sep 24. J Exp Bot. 2013. PMID: 24064926 Free PMC article.
-
Photosynthesis in a Changing Global Climate: Scaling Up and Scaling Down in Crops.Front Plant Sci. 2020 Jul 6;11:882. doi: 10.3389/fpls.2020.00882. eCollection 2020. Front Plant Sci. 2020. PMID: 32733499 Free PMC article. Review.
-
Diethyl Aminoethyl Hexanoate Priming Ameliorates Seed Germination via Involvement in Hormonal Changes, Osmotic Adjustment, and Dehydrins Accumulation in White Clover Under Drought Stress.Front Plant Sci. 2021 Jul 29;12:709187. doi: 10.3389/fpls.2021.709187. eCollection 2021. Front Plant Sci. 2021. PMID: 34394164 Free PMC article.
-
An Overview of Signaling Regulons During Cold Stress Tolerance in Plants.Curr Genomics. 2017 Dec;18(6):498-511. doi: 10.2174/1389202918666170228141345. Curr Genomics. 2017. PMID: 29204079 Free PMC article. Review.
References
-
- Aggarwal KK, Saluja D, Sachar RC. Phosphorylation of rubisco in Cicer arietinum: non-phosphoprotein nature of rubisco in Nicotiana tabacum. Phytochemistry. 1993;34:329–335.
-
- Alsheikh MK, Svensson JT, Randall SK. Phosphorylation regulated ion-binding is a property shared by the acidic subclass dehydrins. Plant, Cell and Environment. 2005;28:1114–1122.
-
- Argueso CT, Ferreira FJ, Kieber JJ. Environmental perception avenues: the interaction of cytokinin and environmental response pathways. Plant, Cell and Environment. 2009;32:1147–1160. - PubMed
-
- Bae MS, Cho EJ, Choi E, Park OK. Analysis of the Arabidopsis nuclear proteome and its response to cold stress. The Plant Journal. 2003;36:652–663. - PubMed

