Computational prediction of type III and IV secreted effectors in gram-negative bacteria
- PMID: 20974833
- PMCID: PMC3019878
- DOI: 10.1128/IAI.00537-10
Computational prediction of type III and IV secreted effectors in gram-negative bacteria
Abstract
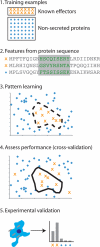
In this review, we provide an overview of the methods employed in four recent studies that described novel methods for computational prediction of secreted effectors from type III and IV secretion systems in Gram-negative bacteria. We present the results of these studies in terms of performance at accurately predicting secreted effectors and similarities found between secretion signals that may reflect biologically relevant features for recognition. We discuss the Web-based tools for secreted effector prediction described in these studies and announce the availability of our tool, the SIEVE server (http://www.sysbep.org/sieve). Finally, we assess the accuracies of the three type III effector prediction methods on a small set of proteins not known prior to the development of these tools that we recently discovered and validated using both experimental and computational approaches. Our comparison shows that all methods use similar approaches and, in general, arrive at similar conclusions. We discuss the possibility of an order-dependent motif in the secretion signal, which was a point of disagreement in the studies. Our results show that there may be classes of effectors in which the signal has a loosely defined motif and others in which secretion is dependent only on compositional biases. Computational prediction of secreted effectors from protein sequences represents an important step toward better understanding the interaction between pathogens and hosts.
Figures
References
-
- Akeda, Y., and J. E. Galan. 2005. Chaperone release and unfolding of substrates in type III secretion. Nature 437:911-915. - PubMed
-
- Amor, J. C., J. Swails, X. Zhu, C. R. Roy, H. Nagai, A. Ingmundson, X. Cheng, and R. A. Kahn. 2005. The structure of RalF, an ADP-ribosylation factor guanine nucleotide exchange factor from Legionella pneumophila, reveals the presence of a cap over the active site. J. Biol. Chem. 280:1392-1400. - PubMed
-
- Anderson, D. M., and O. Schneewind. 1997. A mRNA signal for the type III secretion of Yop proteins by Yersinia enterocolitica. Science 278:1140-1143. - PubMed
-
- Bailey, L., A. Gylfe, C. Sundin, S. Muschiol, M. Elofsson, P. Nordstrom, B. Henriques-Normark, R. Lugert, A. Waldenstrom, H. Wolf-Watz, and S. Bergstrom. 2007. Small molecule inhibitors of type III secretion in Yersinia block the Chlamydia pneumoniae infection cycle. FEBS Lett. 581:587-595. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

