Chronic hypoxia impairs muscle function in the Drosophila model of Duchenne's muscular dystrophy (DMD)
- PMID: 20975992
- PMCID: PMC2958114
- DOI: 10.1371/journal.pone.0013450
Chronic hypoxia impairs muscle function in the Drosophila model of Duchenne's muscular dystrophy (DMD)
Abstract
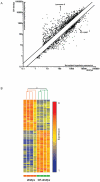
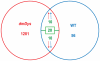
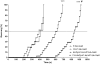
Duchenne's muscular dystrophy (DMD) is a severe progressive myopathy caused by mutations in the DMD gene leading to a deficiency of the dystrophin protein. Due to ongoing muscle necrosis in respiratory muscles late-stage DMD is associated with respiratory insufficiency and chronic hypoxia (CH). To understand the effects of CH on dystrophin-deficient muscle in vivo, we exposed the Drosophila model for DMD (dmDys) to CH during a 16-day ascent to the summit of Mount Denali/McKinley (6194 meters above sea level). Additionally, dmDys and wild type (WT) flies were also exposed to CH in laboratory simulations of high altitude hypoxia. Expression profiling was performed using Affymetrix GeneChips® and validated using qPCR. Hypoxic dmDys differentially expressed 1281 genes, whereas the hypoxic WT flies differentially expressed 56 genes. Interestingly, a number of genes (e.g. heat shock proteins) were discordantly regulated in response to CH between dmDys and WT. We tested the possibility that the disparate molecular responses of dystrophin-deficient tissues to CH could adversely affect muscle by performing functional assays in vivo. Normoxic and CH WT and dmDys flies were challenged with acute hypoxia and time-to-recover determined as well as subjected to climbing tests. Impaired performance was noted for CH-dmDys compared to normoxic dmDys or WT flies (rank order: Normoxic-WT ≈ CH-WT> Normoxic-dmDys> CH-dmDys). These data suggest that dystrophin-deficiency is associated with a disparate, pathological hypoxic stress response(s) and is more sensitive to hypoxia induced muscle dysfunction in vivo. We hypothesize that targeting/correcting the disparate molecular response(s) to hypoxia may offer a novel therapeutic strategy in DMD.
Conflict of interest statement
Figures


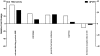
References
-
- Engel AG, Franzini-Armstrong C. New York: McGraw-Hill; 1994. Myology.
-
- Hoffman EP, Brown RH, Kunkel LM. Dystrophin: the protein product of the Duchene muscular dystrophy locus. Cell. 1987;51:919–928. - PubMed
-
- Koenig M, Monaco AP, Kunkel LM. The complete sequence of dystrophin predicts a rod-shaped cytoskeletal protein. Cell. 1988;53:219–226. - PubMed
-
- Bach JR, Ishikawa Y, Kim H. Prevention of pulmonary morbidity for patients with Duchenne muscular dystrophy. Chest. 1997;112:1024–1028. - PubMed
-
- Inkley SR, Oldenburg FC, Vignos PJ., Jr Pulmonary function in Duchenne muscular dystrophy related to stage of disease. Am J Med. 1974;56:297–306. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

