Discovery of recurrent t(6;7)(p25.3;q32.3) translocations in ALK-negative anaplastic large cell lymphomas by massively parallel genomic sequencing
- PMID: 21030553
- PMCID: PMC3035081
- DOI: 10.1182/blood-2010-08-303305
Discovery of recurrent t(6;7)(p25.3;q32.3) translocations in ALK-negative anaplastic large cell lymphomas by massively parallel genomic sequencing
Abstract
The genetics of peripheral T-cell lymphomas are poorly understood. The most well-characterized abnormalities are translocations involving ALK, occurring in approximately half of anaplastic large cell lymphomas (ALCLs). To gain insight into the genetics of ALCLs lacking ALK translocations, we combined mate-pair DNA library construction, massively parallel ("Next Generation") sequencing, and a novel bioinformatic algorithm. We identified a balanced translocation disrupting the DUSP22 phosphatase gene on 6p25.3 and adjoining the FRA7H fragile site on 7q32.3 in a systemic ALK-negative ALCL. Using fluorescence in situ hybridization, we demonstrated that the t(6;7)(p25.3;q32.3) was recurrent in ALK-negative ALCLs. Furthermore, t(6;7)(p25.3;q32.3) was associated with down-regulation of DUSP22 and up-regulation of MIR29 microRNAs on 7q32.3. These findings represent the first recurrent translocation reported in ALK-negative ALCL and highlight the utility of massively parallel genomic sequencing to discover novel translocations in lymphoma and other cancers.
Figures
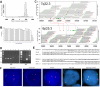

References
-
- Rowley JD. Chromosomal translocations: revisited yet again. Blood. 2008;112(6):2183–2189. - PubMed
-
- Swerdlow S, Campo E, Harris N, et al., editors. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. 4th ed. Lyon, France: International Agency for Research on Cancer; 2008.
-
- Gascoyne RD, Aoun P, Wu D, et al. Prognostic significance of anaplastic lymphoma kinase (ALK) protein expression in adults with anaplastic large cell lymphoma. Blood. 1999;93(11):3913–3921. - PubMed
-
- Li R, Morris SW. Development of anaplastic lymphoma kinase (ALK) small-molecule inhibitors for cancer therapy. Med Res Rev. 2008;28(3):372–412. - PubMed
-
- Mason DY, Harris NL, Delsol G, et al. Anaplastic large cell lymphoma, ALK-negative. In: Swerdlow S, Campo E, Harris N, et al., editors. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. 4th ed. Lyon, France: International Agency for Research on Cancer; 2008. pp. 317–319.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

