Regulating DnaA complex assembly: it is time to fill the gaps
- PMID: 21035377
- PMCID: PMC3005629
- DOI: 10.1016/j.mib.2010.10.001
Regulating DnaA complex assembly: it is time to fill the gaps
Abstract
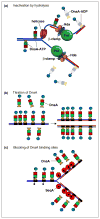
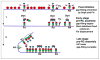
New rounds of bacterial chromosome replication are triggered during each cell division cycle by the initiator protein, DnaA. For precise timing, interactions of DnaA-ATP monomers with the replication origin, oriC, must be carefully regulated during formation of complexes that unwind origin DNA and load replicative helicase. Recent studies in Escherichia coli suggest that high and low affinity DnaA recognition sites are positioned within oriC to direct staged assembly of bacterial pre-replication complexes, with DnaA contacting low affinity sites as it oligomerizes to 'fill the gaps' between high affinity sites. The wide variability of oriC DnaA recognition site patterns seen in nature may reflect myriad gap-filling strategies needed to couple oriC function to the lifestyle of different bacterial types.
Copyright © 2010 Elsevier Ltd. All rights reserved.
Figures

References
-
- Fuller RS, Funnell BE, Kornberg A. The dnaA protein complex with the E. coli chromosomal replication origin (oriC) and other DNA sites. Cell. 1984;38:889–900. - PubMed
-
- Crooke E, Thresher R, Hwang DS, Griffith J, Kornberg A. Replicatively active complexes of DnaA protein and the Escherichia coli chromosomal origin observed in the electron microscope. J Mol Biol. 1993;233:16–24. - PubMed
-
- Margulies C, Kaguni JM. Ordered and sequential binding of DnaA protein to oriC, the chromosomal origin of Escherichia coli. J Biol Chem. 1996;271:17035–17040. - PubMed
-
- Kawakami H, Katayama T. DnaA, ORC, and Cdc6: similarity beyond the domains of life and diversity. Biochem Cell Biol. 2010;88:49–62. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

