A discontinuous DNA glycosylase domain in a family of enzymes that excise 5-methylcytosine
- PMID: 21036872
- PMCID: PMC3045615
- DOI: 10.1093/nar/gkq982
A discontinuous DNA glycosylase domain in a family of enzymes that excise 5-methylcytosine
Abstract
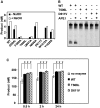
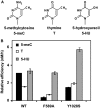
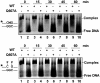
DNA cytosine methylation (5-meC) is a widespread epigenetic mark associated to gene silencing. In plants, DEMETER-LIKE (DML) proteins typified by Arabidopsis REPRESSOR OF SILENCING 1 (ROS1) initiate active DNA demethylation by catalyzing 5-meC excision. DML proteins belong to the HhH-GPD superfamily, the largest and most functionally diverse group of DNA glycosylases, but the molecular properties that underlie their capacity to specifically recognize and excise 5-meC are largely unknown. We have found that sequence similarity to HhH-GPD enzymes in DML proteins is actually distributed over two non-contiguous segments connected by a predicted disordered region. We used homology-based modeling to locate candidate residues important for ROS1 function in both segments, and tested our predictions by site-specific mutagenesis. We found that amino acids T606 and D611 are essential for ROS1 DNA glycosylase activity, whereas mutations in either of two aromatic residues (F589 and Y1028) reverse the characteristic ROS1 preference for 5-meC over T. We also found evidence suggesting that ROS1 uses Q607 to flip out 5-meC, while the contiguous N608 residue contributes to sequence-context specificity. In addition to providing novel insights into the molecular basis of 5-meC excision, our results reveal that ROS1 and its DML homologs possess a discontinuous catalytic domain that is unprecedented among known DNA glycosylases.
Figures



Similar articles
-
Active DNA Demethylation in Plants.Int J Mol Sci. 2019 Sep 21;20(19):4683. doi: 10.3390/ijms20194683. Int J Mol Sci. 2019. PMID: 31546611 Free PMC article. Review.
-
Early steps of active DNA demethylation initiated by ROS1 glycosylase require three putative helix-invading residues.Nucleic Acids Res. 2013 Oct;41(18):8654-64. doi: 10.1093/nar/gkt625. Epub 2013 Jul 18. Nucleic Acids Res. 2013. PMID: 23868090 Free PMC article.
-
DEMETER and REPRESSOR OF SILENCING 1 encode 5-methylcytosine DNA glycosylases.Proc Natl Acad Sci U S A. 2006 May 2;103(18):6853-8. doi: 10.1073/pnas.0601109103. Epub 2006 Apr 19. Proc Natl Acad Sci U S A. 2006. PMID: 16624880 Free PMC article.
-
Methylation-independent DNA binding modulates specificity of Repressor of Silencing 1 (ROS1) and facilitates demethylation in long substrates.J Biol Chem. 2010 Jul 23;285(30):23032-9. doi: 10.1074/jbc.M110.124578. Epub 2010 May 19. J Biol Chem. 2010. PMID: 20489198 Free PMC article.
-
Preventing transcriptional gene silencing by active DNA demethylation.FEBS Lett. 2005 Oct 31;579(26):5889-98. doi: 10.1016/j.febslet.2005.08.039. Epub 2005 Aug 31. FEBS Lett. 2005. PMID: 16162337 Review.
Cited by
-
The recognition domain of the methyl-specific endonuclease McrBC flips out 5-methylcytosine.Nucleic Acids Res. 2012 Aug;40(15):7552-62. doi: 10.1093/nar/gks332. Epub 2012 May 8. Nucleic Acids Res. 2012. PMID: 22570415 Free PMC article.
-
Recent advances in the structural mechanisms of DNA glycosylases.Biochim Biophys Acta. 2013 Jan;1834(1):247-71. doi: 10.1016/j.bbapap.2012.10.005. Epub 2012 Oct 14. Biochim Biophys Acta. 2013. PMID: 23076011 Free PMC article. Review.
-
A DNA 3' phosphatase functions in active DNA demethylation in Arabidopsis.Mol Cell. 2012 Feb 10;45(3):357-70. doi: 10.1016/j.molcel.2011.11.034. Mol Cell. 2012. PMID: 22325353 Free PMC article.
-
Active DNA Demethylation in Plants.Int J Mol Sci. 2019 Sep 21;20(19):4683. doi: 10.3390/ijms20194683. Int J Mol Sci. 2019. PMID: 31546611 Free PMC article. Review.
-
The Mechanisms of Generation, Recognition, and Erasure of DNA 5-Methylcytosine and Thymine Oxidations.J Biol Chem. 2015 Aug 21;290(34):20723-20733. doi: 10.1074/jbc.R115.656884. Epub 2015 Jul 7. J Biol Chem. 2015. PMID: 26152719 Free PMC article. Review.
References
-
- Zemach A, McDaniel IE, Silva P, Zilberman D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science. 2010;328:916–919. - PubMed
-
- Roldan-Arjona T, Ariza RR. DNA demethylation. In: Grosjean H, editor. DNA and RNA modification Enzymes: Comparative Structure, Mechanism, Functions, Cellular Interactions and Evolution. Austin, TX: Landes Bioscience; 2009. pp. 149–161.
-
- Robertson KD. DNA methylation and human disease. Nat. Rev. Genet. 2005;6:597–610. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

