Cut-like homeobox 1 (CUX1) regulates expression of the fat mass and obesity-associated and retinitis pigmentosa GTPase regulator-interacting protein-1-like (RPGRIP1L) genes and coordinates leptin receptor signaling
- PMID: 21037323
- PMCID: PMC3023512
- DOI: 10.1074/jbc.M110.188482
Cut-like homeobox 1 (CUX1) regulates expression of the fat mass and obesity-associated and retinitis pigmentosa GTPase regulator-interacting protein-1-like (RPGRIP1L) genes and coordinates leptin receptor signaling
Abstract
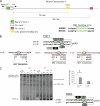
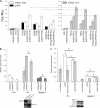
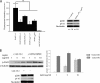
The first intron of FTO contains common single nucleotide polymorphisms associated with body weight and adiposity in humans. In an effort to identify the molecular basis for this association, we discovered that FTO and RPGRIP1L (a ciliary gene located in close proximity to the transcriptional start site of FTO) are regulated by isoforms P200 and P110 of the transcription factor, CUX1. This regulation occurs via a single AATAAATA regulatory site (conserved in the mouse) within the FTO intronic region associated with adiposity in humans. Single nucleotide polymorphism rs8050136 (located in this regulatory site) affects binding affinities of P200 and P110. Promoter-probe analysis revealed that binding of P200 to this site represses FTO, whereas binding of P110 increases transcriptional activity from the FTO as well as RPGRIP1L minimal promoters. Reduced expression of Fto or Rpgrip1l affects leptin receptor isoform b trafficking and leptin signaling in N41 mouse hypothalamic or N2a neuroblastoma cells in vitro. Leptin receptor clusters in the vicinity of the cilium of arcuate hypothalamic neurons in C57BL/6J mice treated with leptin, but not in fasted mice, suggesting a potentially important role of the cilium in leptin signaling that is, in part, regulated by FTO and RPGRIP1L. Decreased Fto/Rpgrip1l expression in the arcuate hypothalamus coincides with decreased nuclear enzymatic activity of a protease (cathepsin L) that has been shown to cleave full-length CUX1 (P200) to P110. P200 disrupts (whereas P110 promotes) leptin receptor isoform b clustering in the vicinity of the cilium in vitro. Clustering of the receptor coincides with increased leptin signaling as reflected in protein levels of phosphorylated Stat3 (p-Stat3). Association of the FTO locus with adiposity in humans may reflect functional consequences of A/C alleles at rs8050136. The obesity-risk (A) allele shows reduced affinity for the FTO and RPGRIP1L transcriptional activator P110, leading to the following: 1) decreased FTO and RPGRIP1L mRNA levels; 2) reduced LEPR trafficking to the cilium; and, as a consequence, 3) a diminished cellular response to leptin.
Figures






References
-
- Frayling T. M., Timpson N. J., Weedon M. N., Zeggini E., Freathy R. M., Lindgren C. M., Perry J. R., Elliott K. S., Lango H., Rayner N. W., Shields B., Harries L. W., Barrett J. C., Ellard S., Groves C. J., Knight B., Patch A. M., Ness A. R., Ebrahim S., Lawlor D. A., Ring S. M., Ben-Shlomo Y., Jarvelin M. R., Sovio U., Bennett A. J., Melzer D., Ferrucci L., Loos R. J., Barroso I., Wareham N. J., Karpe F., Owen K. R., Cardon L. R., Walker M., Hitman G. A., Palmer C. N., Doney A. S., Morris A. D., Smith G. D., Hattersley A. T., McCarthy M. I. (2007) Science 316, 889–894 - PMC - PubMed
-
- Scuteri A., Sanna S., Chen W. M., Uda M., Albai G., Strait J., Najjar S., Nagaraja R., Orrú M., Usala G., Dei M., Lai S., Maschio A., Busonero F., Mulas A., Ehret G. B., Fink A. A., Weder A. B., Cooper R. S., Galan P., Chakravarti A., Schlessinger D., Cao A., Lakatta E., Abecasis G. R. (2007) PLoS Genet. 3, e115 - PMC - PubMed
-
- Meyre D., Delplanque J., Chèvre J. C., Lecoeur C., Lobbens S., Gallina S., Durand E., Vatin V., Degraeve F., Proença C., Gaget S., Körner A., Kovacs P., Kiess W., Tichet J., Marre M., Hartikainen A. L., Horber F., Potoczna N., Hercberg S., Levy-Marchal C., Pattou F., Heude B., Tauber M., McCarthy M. I., Blakemore A. I., Montpetit A., Polychronakos C., Weill J., Coin L. J., Asher J., Elliott P., Järvelin M. R., Visvikis-Siest S., Balkau B., Sladek R., Balding D., Walley A., Dina C., Froguel P. (2009) Nat. Genet. 41, 157–159 - PubMed
-
- Thorleifsson G., Walters G. B., Gudbjartsson D. F., Steinthorsdottir V., Sulem P., Helgadottir A., Styrkarsdottir U., Gretarsdottir S., Thorlacius S., Jonsdottir I., Jonsdottir T., Olafsdottir E. J., Olafsdottir G. H., Jonsson T., Jonsson F., Borch-Johnsen K., Hansen T., Andersen G., Jorgensen T., Lauritzen T., Aben K. K., Verbeek A. L., Roeleveld N., Kampman E., Yanek L. R., Becker L. C., Tryggvadottir L., Rafnar T., Becker D. M., Gulcher J., Kiemeney L. A., Pedersen O., Kong A., Thorsteinsdottir U., Stefansson K. (2009) Nat. Genet. 41, 18–24 - PubMed
-
- Gerken T., Girard C. A., Tung Y. C., Webby C. J., Saudek V., Hewitson K. S., Yeo G. S., McDonough M. A., Cunliffe S., McNeill L. A., Galvanovskis J., Rorsman P., Robins P., Prieur X., Coll A. P., Ma M., Jovanovic Z., Farooqi I. S., Sedgwick B., Barroso I., Lindahl T., Ponting C. P., Ashcroft F. M., O'Rahilly S., Schofield C. J. (2007) Science 318, 1469–1472 - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

