Kinesin 3 and cytoplasmic dynein mediate interkinetic nuclear migration in neural stem cells
- PMID: 21037580
- PMCID: PMC3059207
- DOI: 10.1038/nn.2665
Kinesin 3 and cytoplasmic dynein mediate interkinetic nuclear migration in neural stem cells
Abstract
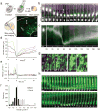
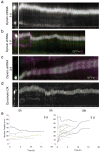
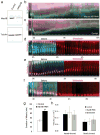
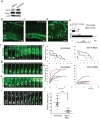
Radial glial progenitor cells exhibit bidirectional cell cycle-dependent nuclear oscillations. The purpose and underlying mechanism of this unusual 'interkinetic nuclear migration' are poorly understood. We investigated the basis for this behavior by live imaging of nuclei, centrosomes and microtubules in embryonic rat brain slices, coupled with the use of RNA interference (RNAi) and the myosin inhibitor blebbistatin. We found that nuclei migrated independent of centrosomes and unidirectionally away from or toward the ventricular surface along microtubules, which were uniformly oriented from the ventricular surface to the pial surface of the brain. RNAi directed against cytoplasmic dynein specifically inhibited nuclear movement toward the apical surface. An RNAi screen of kinesin genes identified Kif1a, a member of the kinesin-3 family, as the motor for basally directed nuclear movement. These observations provide direct evidence that kinesins are involved in nuclear migration and neurogenesis and suggest that a cell cycle-dependent switch between distinct microtubule motors drives interkinetic nuclear migration.
Figures



Comment in
-
Interkinetic nuclear migration: reciprocal activities of dynein and kinesin.Cell Adh Migr. 2011 Jul-Aug;5(4):277-9. doi: 10.4161/cam.5.4.17432. Epub 2011 Jul 1. Cell Adh Migr. 2011. PMID: 21836392 Free PMC article.
Similar articles
-
Regulation of interkinetic nuclear migration by cell cycle-coupled active and passive mechanisms in the developing brain.EMBO J. 2011 May 4;30(9):1690-704. doi: 10.1038/emboj.2011.81. Epub 2011 Mar 25. EMBO J. 2011. PMID: 21441895 Free PMC article.
-
Apical movement during interkinetic nuclear migration is a two-step process.Dev Biol. 2012 Oct 1;370(1):33-41. doi: 10.1016/j.ydbio.2012.06.031. Epub 2012 Aug 4. Dev Biol. 2012. PMID: 22884563 Free PMC article.
-
Kinesin-1, -2, and -3 motors use family-specific mechanochemical strategies to effectively compete with dynein during bidirectional transport.Elife. 2022 Sep 20;11:e82228. doi: 10.7554/eLife.82228. Elife. 2022. PMID: 36125250 Free PMC article.
-
Emerging roles for motor proteins in progenitor cell behavior and neuronal migration during brain development.Cytoskeleton (Hoboken). 2016 Oct;73(10):566-576. doi: 10.1002/cm.21293. Epub 2016 Apr 4. Cytoskeleton (Hoboken). 2016. PMID: 26994401 Review.
-
Interkinetic nuclear migration: a mysterious process in search of a function.Dev Growth Differ. 2012 Apr;54(3):306-16. doi: 10.1111/j.1440-169X.2012.01342.x. Dev Growth Differ. 2012. PMID: 22524603 Free PMC article. Review.
Cited by
-
The nuclear lamins: flexibility in function.Nat Rev Mol Cell Biol. 2013 Jan;14(1):13-24. doi: 10.1038/nrm3488. Epub 2012 Dec 5. Nat Rev Mol Cell Biol. 2013. PMID: 23212477 Review.
-
Multiple modes of cytoplasmic dynein regulation.Nat Cell Biol. 2012 Feb 29;14(3):224-30. doi: 10.1038/ncb2420. Nat Cell Biol. 2012. PMID: 22373868
-
Mechanical principles of nuclear shaping and positioning.J Cell Biol. 2018 Oct 1;217(10):3330-3342. doi: 10.1083/jcb.201804052. Epub 2018 Sep 7. J Cell Biol. 2018. PMID: 30194270 Free PMC article. Review.
-
Otic Neurogenesis in Xenopus laevis: Proliferation, Differentiation, and the Role of Eya1.Front Neuroanat. 2021 Sep 20;15:722374. doi: 10.3389/fnana.2021.722374. eCollection 2021. Front Neuroanat. 2021. PMID: 34616280 Free PMC article.
-
Live-cell imaging: new avenues to investigate retinal regeneration.Neural Regen Res. 2017 Aug;12(8):1210-1219. doi: 10.4103/1673-5374.213533. Neural Regen Res. 2017. PMID: 28966629 Free PMC article. Review.
References
-
- Solecki DJ, Model L, Gaetz J, Kapoor TM, Hatten ME. Par6alpha signaling controls glial-guided neuronal migration. Nat Neurosci. 2004;7:1195–1203. - PubMed
-
- Tsai JW, Bremner KH, Vallee RB. Dual subcellular roles for LIS1 and dynein in radial neuronal migration in live brain tissue. Nat Neurosci. 2007;10:970–979. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

