Alternative pre-mRNA splicing regulation in cancer: pathways and programs unhinged
- PMID: 21041405
- PMCID: PMC2964746
- DOI: 10.1101/gad.1973010
Alternative pre-mRNA splicing regulation in cancer: pathways and programs unhinged
Abstract
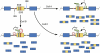
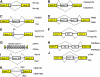
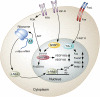
Alternative splicing of mRNA precursors is a nearly ubiquitous and extremely flexible point of gene control in humans. It provides cells with the opportunity to create protein isoforms of differing, even opposing, functions from a single gene. Cancer cells often take advantage of this flexibility to produce proteins that promote growth and survival. Many of the isoforms produced in this manner are developmentally regulated and are preferentially re-expressed in tumors. Emerging insights into this process indicate that pathways that are frequently deregulated in cancer often play important roles in promoting aberrant splicing, which in turn contributes to all aspects of tumor biology.
Figures

References
-
- Arch R, Wirth K, Hofmann M, Ponta H, Matzku S, Herrlich P, Zoller M 1992. Participation in normal immune responses of a metastasis-inducing splice variant of CD44. Science 257: 682–685 - PubMed
-
- Babic I, Cherry E, Fujita DJ 2006. SUMO modification of Sam68 enhances its ability to repress cyclin D1 expression and inhibits its ability to induce apoptosis. Oncogene 25: 4955–4964 - PubMed
-
- Barash Y, Calarco JA, Gao W, Pan Q, Wang X, Shai O, Blencowe BJ, Frey BJ 2010. Deciphering the splicing code. Nature 465: 53–59 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
