The spatial-functional coupling of box C/D and C'/D' RNPs is an evolutionarily conserved feature of the eukaryotic box C/D snoRNP nucleotide modification complex
- PMID: 21041475
- PMCID: PMC3019978
- DOI: 10.1128/MCB.00918-10
The spatial-functional coupling of box C/D and C'/D' RNPs is an evolutionarily conserved feature of the eukaryotic box C/D snoRNP nucleotide modification complex
Abstract
Box C/D ribonucleoprotein particles guide the 2'-O-ribose methylation of target nucleotides in both archaeal and eukaryotic RNAs. These complexes contain two functional centers, assembled around the C/D and C'/D' motifs in the box C/D RNA. The C/D and C'/D' RNPs of the archaeal snoRNA-like RNP (sRNP) are spatially and functionally coupled. Here, we show that similar coupling also occurs in eukaryotic box C/D snoRNPs. The C/D RNP guided 2'-O-methylation when the C'/D' motif was either mutated or ablated. In contrast, the C'/D' RNP was inactive as an independent complex. Additional experiments demonstrated that the internal C'/D' RNP is spatially coupled to the terminal box C/D complex. Pulldown experiments also indicated that all four core proteins are independently recruited to the box C/D and C'/D' motifs. Therefore, the spatial-functional coupling of box C/D and C'/D' RNPs is an evolutionarily conserved feature of both archaeal and eukaryotic box C/D RNP complexes.
Figures
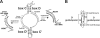
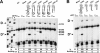
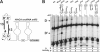


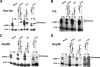
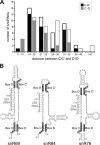
References
-
- Aittaleb, M., R. Rashid, Q. Chen, J. Almer, C. Daniels, and H. Lee. 2003. Structure and function of archaeal box C/D sRNP core proteins. Nat. Struct. Biol. 10:256-263. - PubMed
-
- Cavaillé, J., M. Nicoloso, and J. P. Bachellerie. 1996. Targeted ribose methylation of RNA in vivo directed by tailored antisense RNA guides. Nature 383:732-735. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
