Taming free energy landscapes with RNA chaperones
- PMID: 21045544
- PMCID: PMC3073327
- DOI: 10.4161/rna.7.6.13615
Taming free energy landscapes with RNA chaperones
Abstract
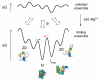
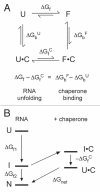
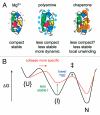
Many non-coding RNAs fold into complex three-dimensional structures, yet the self-assembly of RNA structure is hampered by mispairing, weak tertiary interactions, electrostatic barriers, and the frequent requirement that the 5' and 3' ends of the transcript interact. This rugged free energy landscape for RNA folding means that some RNA molecules in a population rapidly form their native structure, while many others become kinetically trapped in misfolded conformations. Transient binding of RNA chaperone proteins destabilize misfolded intermediates and lower the transition states between conformations, producing a smoother landscape that increases the rate of folding and the probability that a molecule will find the native structure. DEAD-box proteins couple the chemical potential of ATP hydrolysis with repetitive cycles of RNA binding and release, expanding the range of conditions under which they can refold RNA structures.
Figures

References
-
- Herschlag D. RNA chaperones and the RNA folding problem. J Biol Chem. 1995;270:77–79. - PubMed
-
- Thirumalai D, Hyeon C. RNA and protein folding: common themes and variations. Biochemistry. 2005;44:4957–4970. - PubMed
-
- Weeks KM. Protein-facilitated RNA folding. Curr Opin Struct Biol. 1997;7:336–342. - PubMed
-
- Rajkowitsch L, Chen D, Stampfl S, Semrad K, Waldsich C, Mayer O, et al. RNA chaperones, RNA annealers and RNA helicases. RNA Biol. 2007;4:118–130. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
