An atlas of gene regulatory networks reveals multiple three-gene mechanisms for interpreting morphogen gradients
- PMID: 21045819
- PMCID: PMC3010108
- DOI: 10.1038/msb.2010.74
An atlas of gene regulatory networks reveals multiple three-gene mechanisms for interpreting morphogen gradients
Abstract
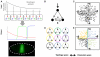
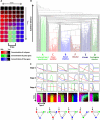
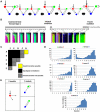
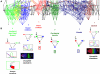
The interpretation of morphogen gradients is a pivotal concept in developmental biology, and several mechanisms have been proposed to explain how gene regulatory networks (GRNs) achieve concentration-dependent responses. However, the number of different mechanisms that may exist for cells to interpret morphogens, and the importance of design features such as feedback or local cell-cell communication, is unclear. A complete understanding of such systems will require going beyond a case-by-case analysis of real morphogen interpretation mechanisms and mapping out a complete GRN 'design space.' Here, we generate a first atlas of design space for GRNs capable of patterning a homogeneous field of cells into discrete gene expression domains by interpreting a fixed morphogen gradient. We uncover multiple very distinct mechanisms distributed discretely across the atlas, thereby expanding the repertoire of morphogen interpretation network motifs. Analyzing this diverse collection of mechanisms also allows us to predict that local cell-cell communication will rarely be responsible for the basic dose-dependent response of morphogen interpretation networks.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

References
-
- Bayly RD, Ngo M, Aglyamova GV, Agarwala S (2007) Regulation of ventral midbrain patterning by Hedgehog signaling. Development 134: 2115–2124 - PubMed
-
- Briscoe J, Chen Y, Jessell TM, Struhl G (2001) A hedgehog-insensitive form of patched provides evidence for direct long-range morphogen activity of sonic hedgehog in the neural tube. Mol Cell 7: 1279–1291 - PubMed
-
- Chouard T (2008) Beneath the surface. Nature 456: 300–303 - PubMed
-
- Clyde DE, Corado MS, Wu X, Paré A, Papatsenko D, Small S (2003) A self-organizing system of repressor gradients establishes segmental complexity in Drosophila. Nature 18: 849–853 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

