Structural evolution of protein-biofilms: Simulations and experiments
- PMID: 21045923
- PMCID: PMC2967234
- DOI: 10.1063/1.3488672
Structural evolution of protein-biofilms: Simulations and experiments
Abstract
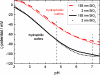
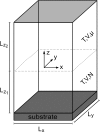
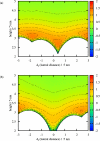
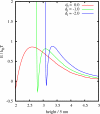
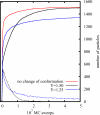
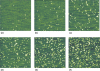
The control of biofilm formation is a challenging goal that has not been reached yet in many aspects. One unsolved question is the role of van der Waals forces and another is the importance of mutual interactions between the adsorbing and the adsorbed biomolecules ("critical crowding"). In this study, a combined experimental and theoretical approach is presented, which fundamentally probes both aspects. On three model proteins-lysozyme, α-amylase, and bovine serum albumin-the adsorption kinetics is studied experimentally. Composite substrates are used enabling a separation of the short- and the long-range forces. Although usually neglected, experimental evidence is given for the influence of van der Waals forces on the protein adsorption as revealed by in situ ellipsometry. The three proteins were chosen for their different conformational stabilities in order to investigate the influence of conformational changes on the adsorption kinetics. Monte Carlo simulations are used to develop a model for these experimental results by assuming an internal degree of freedom to represent conformational changes. The simulations also provide data on the distribution of adsorption sites. By in situ atomic force microscopy we can also test this distribution experimentally, which opens the possibility to, e.g., investigate the interactions between adsorbed proteins.
Figures


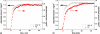


Similar articles
-
Subsurface influence on the structure of protein adsorbates as revealed by in situ X-ray reflectivity.Langmuir. 2012 May 22;28(20):7747-56. doi: 10.1021/la300850g. Epub 2012 May 8. Langmuir. 2012. PMID: 22533829
-
Specific and nonspecific interaction forces between Escherichia coli and silicon nitride, determined by poisson statistical analysis.Langmuir. 2006 Aug 15;22(17):7296-301. doi: 10.1021/la0533415. Langmuir. 2006. PMID: 16893229
-
Incorporating water-release and lateral protein interactions in modeling equilibrium adsorption for ion-exchange chromatography.J Chromatogr A. 2006 Sep 8;1126(1-2):304-10. doi: 10.1016/j.chroma.2006.06.058. Epub 2006 Jul 18. J Chromatogr A. 2006. PMID: 16846610
-
Characterization of van der Waals type bimodal,- lambda,- meta- and spinodal phase transitions in liquid mixtures, solid suspensions and thin films.Adv Colloid Interface Sci. 2018 Mar;253:66-116. doi: 10.1016/j.cis.2018.01.002. Adv Colloid Interface Sci. 2018. PMID: 29422417 Review.
-
Experimental techniques to study protein-surfactant interactions: New insights into competitive adsorptions via drop subphase and interface exchange.Adv Colloid Interface Sci. 2022 Mar;301:102601. doi: 10.1016/j.cis.2022.102601. Epub 2022 Jan 13. Adv Colloid Interface Sci. 2022. PMID: 35114446 Review.
Cited by
-
Hydroxyapatite Pellets as Versatile Model Surfaces for Systematic Adhesion Studies on Enamel: A Force Spectroscopy Case Study.ACS Biomater Sci Eng. 2022 Apr 11;8(4):1476-1485. doi: 10.1021/acsbiomaterials.1c00925. Epub 2022 Mar 9. ACS Biomater Sci Eng. 2022. PMID: 35263544 Free PMC article.
-
Life under flow: A novel microfluidic device for the assessment of anti-biofilm technologies.Biomicrofluidics. 2013 Dec 23;7(6):64118. doi: 10.1063/1.4850796. eCollection 2013 Dec 23. Biomicrofluidics. 2013. PMID: 24454610 Free PMC article.
-
Organosilane deposition for microfluidic applications.Biomicrofluidics. 2011 Sep;5(3):36501-365017. doi: 10.1063/1.3625605. Epub 2011 Aug 16. Biomicrofluidics. 2011. PMID: 22662048 Free PMC article.
-
Kinetic mechanisms in morpholino-DNA surface hybridization.J Am Chem Soc. 2011 Aug 3;133(30):11588-96. doi: 10.1021/ja202631b. Epub 2011 Jul 12. J Am Chem Soc. 2011. PMID: 21699181 Free PMC article.
-
Multiprotein interactions during surface adsorption: a molecular dynamics study of lysozyme aggregation at a charged solid surface.J Phys Chem B. 2011 Jul 21;115(28):8891-900. doi: 10.1021/jp1121239. Epub 2011 Jun 24. J Phys Chem B. 2011. PMID: 21671567 Free PMC article.
References
-
- Denis F. A., Hanarp P., Sutherland D. S., Gold J., Mustin C., Rouxhet P., and Dufrêne Y. F., Langmuir LANGD5 18, 819 (2002).10.1021/la011011o - DOI
-
- Biopolymers at Interfaces, edited by Malmsten M. (Dekker, New York, 2003).
-
- Michael K. E., Vernekar V. N., Keselowsky B. G., Meredith J., Latour R. A., and Garcia A. J., Langmuir LANGD5 19, 8033 (2003).10.1021/la034810a - DOI
LinkOut - more resources
Full Text Sources
